Translate this page into:
CRISPR/Cas9 mediated TaRPK1 root architecture gene mutagenesis confers enhanced wheat yield
⁎Corresponding authors. ymichigan@jnu.ac.kr (Seung Hwan Yang), sfiaz@uoh.edu.pk (Muhammad Ramzan Khan) mrkhan@parc.gov.pk (Muhammad Ramzan Khan)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
CRISPR/Cas9 system has emerged as an efficient tool for sustainable crop improvement. Roots are the “principal hidden organ” that has a crucial function in vascular plants. Receptor-like protein kinase 1 (RPK1) has been reported to regulate root architecture system (RAS), abiotic stress, and yield both in Arabidopsis and rice. We employed a CRISPR/Cas9-based system, namely LR-1 and LR-2 constructs having double guided RNAs transformed via agrobacterium for targeted mutagenesis of TaRPK1 genes to alter the root architecture and hence yield in Triticum aestivum. Sequencing confirmed seven CRISPR/Cas9-based mutated T0 lines of LR-1 constructs and six T0 lines of LR-2 constructs, with an overall mutation efficiency of 41.93%. The T0 plants displayed higher monoallelic mutation compared to the diallelic mutation. 37.5% monoallelic mutation at target site 1 within the D genome by gRNA1 was observed by the LR-1 construct. The LR-2 constructs showed a higher monoallelic mutation frequency of 26.67% at target sites 1 and 2 within A, B, and D genomes. The deletions were mainly short, however longer deletions such as 12d, 17d, 19d, and 20d were detected by gRNA2 of LR-1 construct. Transgenic lines revealed significant alteration in morphology and RSA with a significant increase in number of effective tillers, grain weight, root length, root depth, root volume, and root surface area while reduced root diameter, root angle, and spike length, compared to the wild plants. Tillers and total grain weight increased significantly, suggesting edited lines increased grain production by decreasing spike length. The study validates that CRISPR/Cas9 mediated targeted editing of TaRPK1 is a practical approach for modifying RAS and hence yield enhancement in wheat.
Keywords
RPK1 editing
Agrobacterium-mediated transformation
Root architecture
Grain weight
Wheat
1 Introduction
Roots are the “principal hidden organ” that has a crucial function in vascular plants. Roots are an integral component, mediating the water and nutrient uptake required for the growth of the plant (Ober et al., 2021). The root's uptake of nutrients and water modulates the yields of crops which relies on spatial and temporal distribution of roots in soil (Chen et al., 2020). Root system architecture (RSA) includes length, diameter, length density, depth, volume, and surface area, which organize root structure and affect nutrition and water intake (Alahmad et al., 2019, Djanaguiraman et al., 2019). Among several organs of plants studied, drought mainly affects roots because of its direct association with soil. Drought resistance is correlated directly with the diameter of the root, as bulky roots with huge xylem vessels are effective for the nutrients and water acquisition from deep layers of soil during the conditions of rainfall (Ahmar et al., 2023). In 40 years, drought-stressed plants have quadrupled, and 12 million hectares of land are lost annually to drought and desertification. Worldwide, 79 % of wheat harvesting regions showed drought-induced grain production fluctuation. (Djanaguiraman et al., 2019). Understanding root contributions and variations may improve production and drought-resistant genotypes (Fradgley et al., 2020).
The root systems of dicotyledons and monocotyledons are structurally distinct. Wheat's root system has seminal roots (SR) and crown roots (CR), which grow from shoot basal nodes (Nehe et al., 2021). The seminal roots infiltrate firstly into the soil and stay functional throughout the plant lifecycle (Rufo et al., 2020). These function altogether for the nutrients and water intake from the soil. Bread wheat (Triticum aestivum) is one of the major grain crops with a production and area of 771.71 million tons and 218.54 million hectares, respectively (FAOSTAT. 2017). Due to hexaploidy and an abundance of repetitive and transposable elements, bread wheat has one of the biggest agricultural plant genomes (16 Gb), making genetics, genomics, and breeding difficult (Schilling et al., 2020). Globally, wheat is an essential crop, and more than a quarter of the population relies on it for feed (Gabay et al., 2023). Since wheat is the main staple crop, production needs to be enhanced by 2050 by approximately 38 % to feed the rising population (Djanaguiraman et al., 2019). Drought is one of the primary effects of climate change on agricultural output and growth worldwide. Globally, approximately 79 % displayed variability in grain yield in the harvesting region of wheat due to the fluctuations in temperature and precipitation. Therefore, improvement in root architecture traits, drought tolerance, and high-yield maintenance is a major challenge in the improvement of wheat (Djanaguiraman et al., 2019).
Genome editing is the method of creating specific alterations to a known sequence of DNA. Deletions, insertions, or changes in the gene sequence may modify the protein produced (Brandt et al., 2020). Sequence-specific nucleases (SSNs) like mega-nucleases, zinc-finger nucleases (ZFN), and transcription activator-like effector nuclease (TALENs) methods of editing genomes were applied broadly in plants. However, recently, Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) with Cas nucleases have overtaken these SSNs and mediate the targeting of DNA through guide RNAs (gRNA), which is easy to engineer (Malzahn et al., 2019). CRISPR/Cas9 is considered the most prevalent technology for editing genomes globally, due to its simplicity, versatility, specificity, and flexibility (Li et al., 2019, Ahmar and Gruszka, 2023). It has been used in numerous plant species including Arabidopsis and Nicotiana model-plants, and crop species like maize, rice, and wheat (Uzair et al., 2021, Ahmar et al., 2023).
RPK1 is a Receptor-like protein kinase 1, belongs to the family of LRR-RLKs, comprises six LRR extracellular motifs, a transmembrane domain (TM), and a single cytoplasmic conserved kinase domain in rice (Rahim et al., 2022). Studies have indicated that OsRPK1 negatively modulates polar auxin transport (PAT) and auxin accretion in roots, affecting root architecture (Zou et al., 2014). Suppression of RPK1 caused the growth and increased plant height and tiller numbers. Enhanced expression caused a reduction in roots apical meristem and immature adventitious and lateral roots (Zou et al., 2014). AtRPK1 inhibition, in Arabidopsis thaliana exhibited enhanced salt tolerance whereas up-regulated plants displayed lower degrees of salt tolerance in comparison to the normal plants (Shi et al., 2014). AtRPK1 positively modulates the expression of the CaM1 gene which further modulates ROS production, senescence of the leaf, and ABA response (Dai et al., 2018).
The CRISPR/Cas9 system has successfully demonstrated precise gene editing in many plant systems (Kim et al., 2023). The first reported use of CRISPR-Cas9 to produce a stably genome-edited wheat plant was the targeted knockout of the Mildew Locus O (Mlo), conferring resistance to the powdery mildew pathogen Blumeria graminis f.sp. tritici (Bgt) (Wang et al., 2014). Since the initial publication, CRISPR-Cas9 has targeted wheat genes of agronomical and basic scientific interest, such as TaGW2 to enhance grain weight (Zhang et al., 2018), TaQsd1 to reduce preharvest sprouting (Abe et al., 2019), CENH3 for haploid plant induction (Lv et al., 2020), Sal1 for drought resistance (Mohr et al., 2022). To date, RPK1 disruption via CRISPR/Cas9 system has not employed for wheat to alter its root architecture and related traits. To address the issue, we synthesized two constructs namely LR-1 and LR-2, with two guide RNAs (gRNAs) each, and developed an In-planta Agrobacterium-mediated CRISPR/Cas9 system, hence generating successfully targeted mutations in target sites of TaRPK1 genes in T0 plants with altered root architecture traits and other agronomic traits. We believe that our study has potential application to produce valuable traits in bread wheat and will aid in the improvement of wheat to guarantee global food security.
2 Material and methods
2.1 Wheat germplasm and growing conditions
The seeds of the hexaploid bread wheat cultivar ‘‘Pakistan 2013″ were obtained from the National Institute of Genomics and Advanced Biotechnology (NIGAB), NARC, Islamabad, Pakistan. The control and transgenic (T0) plants were grown in glasshouse at NIGAB. Green-house conditions for day light period of 16 h with 25 ± 3 °C temperature and night period of 8 h with 15 ± 3 °C temperature were kept. Experiments were carried out on complete randomized design (CRD). Three readings were observed for agronomic parameters. The main steps involved in CRISPR-based targeted mutagenesis are summarized in Supplementary Figure S1.
2.2 Target selection and guide RNAs designing
The sequences of the RPK1 genes in the A, B, and D genomes of wheat were retrieved from “Ensemble Plant” through the BLAST of RPK1 sequences of Arabidopsis thaliana and Oryza sativa (Shi et al., 2014, Zou et al., 2014). The sequence of RPK1 with a significant percentage ID and low E-value was selected. The gRNAs were designed manually from exon region, and the possible off-target sites were identified using BLAST search (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The CRISPR/Cas9 system provided by Dr. Bing Yang (Genetics, Development and Cell Biology department, Ames, Iowa State University, USA) was followed. The gRNA expression cassettes were synthesized, with BtgZI restriction site, TGTT, and AAAC at the 5́ ends of sense and antisense strand, respectively. Similarly, from the 2nd exon, another four oligos were designed with BsaI restriction site, GTGT, and AAAC at the 5́ ends of sense and antisense strand, respectively (Table 1).
Gene name and gRNA
Guide RNA sequence (5́ to 3́)
TaRPK-gRNA1-F
TGTTCTTGTTGTTCTTGGTGTTGC
TaRPK-gRNA1-R
AAACGCAACACCAAGAACAACAAG
TaRPK-gRNA2-F
GTGTCCTATGCTTTCCTTGCTGAT
TaRPK-gRNA2-R
AAACATCAGCAAGGAAAGCATAGG
TaRPK-gRNA3-F
GTGTATGGATGTGTTTGATGTCAG
TaRPK-gRNA3-R
AAACCTGACATCAAACACATCCAT
2.3 CRISPR/Cas9 based vector construction
CRISPR/Cas9 system was used for TaRPK1 gene disruption, to unveil its role in wheat. The constructs were synthesized following Bing Yang’s method (Char et al., 2019).
2.4 Transformation and screening of edited wheat plants
The seeds of the Pakistan-2013 wheat cultivar were used following the method of in-planta transformation (Abdul et al., 2011). For in-planta transformation, the seeds were first soaked in 70 % v/v solution.Seeds were then washed with 50 % Clorox for 15 min and were allowed to completely dry. Sowing of seeds was done on Murashige and Skoog(MS) solid medium (Murashige and Skoog, 1962).
The overnight grown liquid cultures of A. tumefaciens strain LBA4404 in LB liquid media were centrifuged at 85000 rpm/10 min, then supplemented with 200 µM acetosyringone and mixed gently. For the three-day-old seedlings, the apical meristem part was pierced up to 1 mm depth, and agrobacterium inoculum was injected into it. After three days of co-cultivation, to remove excess bacteria. The seedlings were then shifted to the selection and regeneration medium containing cefotaxime 250μg/mL; BAP 1.5 mg/L, and hygromycin. The transformed seedlings were then placed in a greenhouse under a controlled environment (22/16 °C with day/night 16/8h photoperiod and 70 % humidity) till maturity.
For the detection of gRNA efficiency in wheat plants, a T7 endonuclease I assay was performed (Kim et al., 2018). To confirm the DNA double-strand break, PCR amplicons were gel purified, denatured, reannealed, and treated with T7EI. The PCR amplicons were sequenced to determine the mutation frequency.
2.5 Root architecture traits and morphological parameters determination
The root architecture traits including root length, depth, diameter, volume, surface area, and root orientation were observed in four-week-old T0 transgenic lines and wild-type plants with the help of RhizoVision software (Seethepalli et al., 2021). Morphological data measured included the number of effective tillers, plant height, spike length, and 1000-grain weight at the maturity stage. A minimum of three replicates from T0 transgenic lines and control plants were recorded for each trait.
2.6 Statistical analysis
All the recorded data was statistically analyzed by using Microsoft Excel 2019. Student t-test was performed to determine the significant differences among the samples (n = 3, *= P < 0.05, **=P < 0.01).
3 Results
3.1 Selection of target homeolog of TaRPK1 gene
The RPK1 sequences in the A, B, and D sub-genomes of wheat retrieved by BLAST from “Ensemble Plant”, with the highest percentage ID and low E-value were located on chromosome 2. TraesCS2A02G176500, TraesCS2B02G202900, and TraesCS2D02G183900 wheat sequence Ensemble IDs showed the highest percentage of similarities 86 %, 85.5 %, and 94.1 %, respectively (Supplementary Table S1). The coding sequences were aligned and sgRNAs were synthesized from the conserved region. The sgRNA1 was designed from the conserved region of A and D genomes, sgRNA2 from A, B, and D genomes, and sgRNA3 from the conserved region of A and B genomes (Supplementary Table S2). These sgRNAs were synthesized from the first exon of the A and B genomes, and the second exon of the D genome (Fig. 1).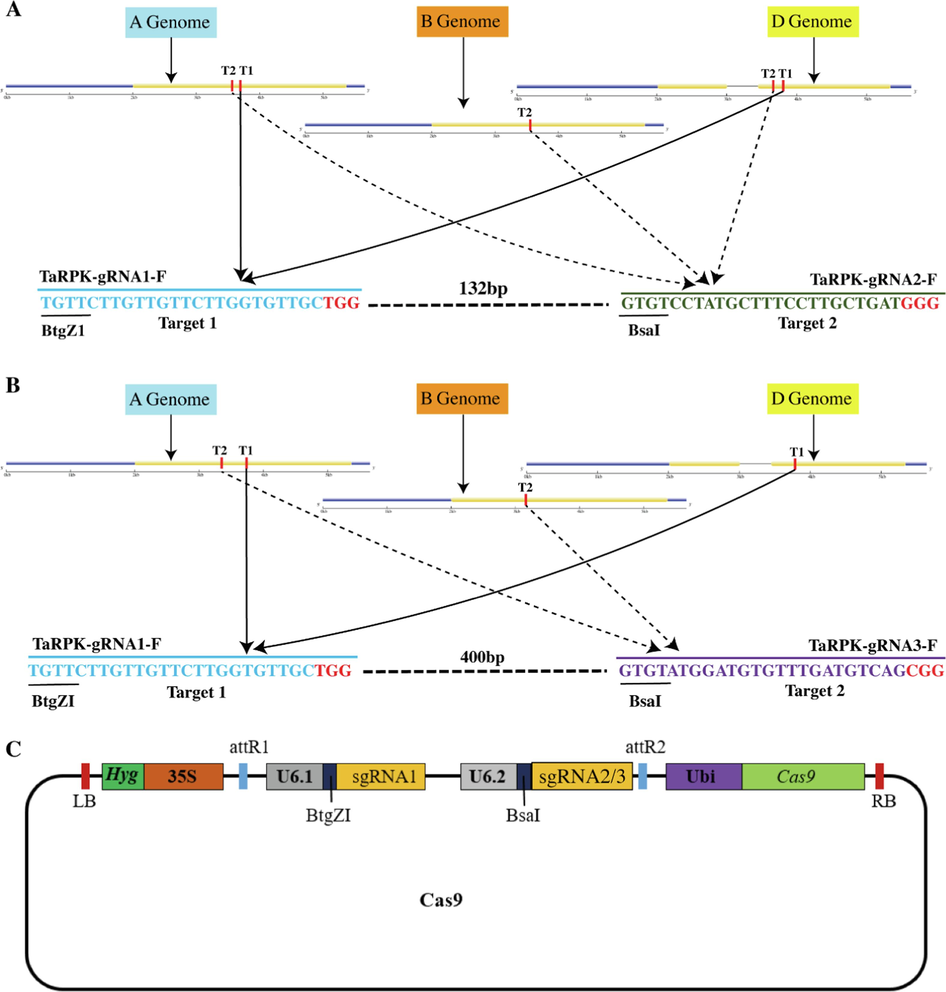
Illustration of sgRNA expression cassette. (A) The representation of TaRPK1-gRNA1 (Target 1) and TaRPK1-gRNA2 (Target 2) for constructing LR1. (B) Indication of TaRPK1-gRNA1 (Target 1) and TaRPK1-gRNA3 (Target 2) for LR2 construct. PAM sites were shown in red color and restriction endonucleases (BtgZI and BsaI) recognition sequences were underlined. (C) Schematic illustration of binary vector transformed into wheat. The constructs TaRPK1-gRNA consists of a cassette with Kanamycin resistant gene, flanked by attR1 and attR2 gateway recombination sequence, inserted into the pOs-Cas9 binary vector via LR recombination reaction. The expression cassette comprises gRNA pairs (TaRPK1-gRNA1 and TaRPK1-gRNA2/3) followed by Cas9 endonuclease regulated by Rice Ubiquitin promoter. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
3.2 CRISPR/Cas9 TaRPK1 vector and in-planta transformation
The binary vector CRISPR/Cas9 containing TaRPK1-sgRNAs were constructed accomplished by gateway recombination reaction (Supplementary Figure S2). The first construct with a combination of sgRNA1 and sgRNA2 is termed LR-1, while the LR-2 construct contains the sgRNA1 and sgRNA3 expression cassettes, respectively. In planta apical meristem transformation being an easy, cheap, and efficient method for transformation was adopted for transformation using CRISPR/cas9 targeted mutagenesis (Fig. 2). More than 25 independent lines were obtained by using this technique.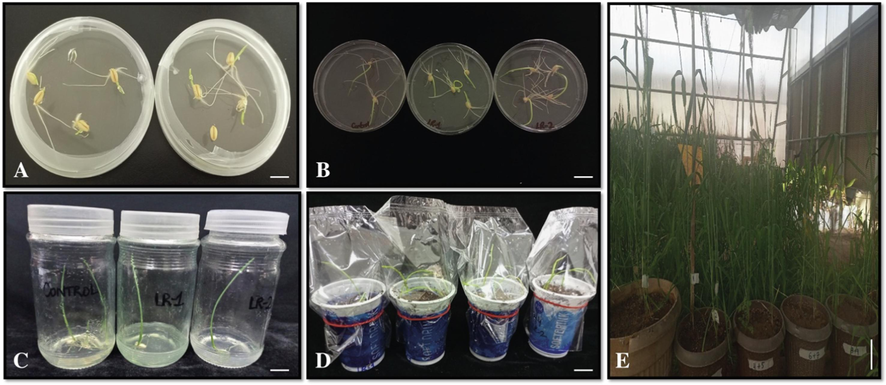
In planta apical meristem transformation CRISPR/Cas9 targeted mutagenesis. (A) Wheat seeds germination on MS medium. (B) Transfer of ex-plant into co-cultivation media. (C) Screening of plants on selection media. Plants with hygromycin resistance survived on selection media. (D) Acclimatization of transplanted seedlings in the growth room, by covering the seedlings with a transparent sheet. (E) Transformed plants shifted to pots in glass house and grown under controlled conditions.
3.3 Mutation detection in T0 plants
The detection of the edit mutation type was performed after the transformation and regeneration of T0 plants. A total of 31 transgenic plants were obtained from screening of selection antibiotic, 16 of which were of LR-1 constructs and 15 of LR-2 constructs (Table 2). Mutation detection in these transgenic plants was performed by genomic DNA extraction from the leaf tissues of T0 plants (Supplementary Figure S3), followed by designing specific primers flanking target sites of A, B, and D genomes (Supplementary Table S3). The PCR products were digested with T7 endonuclease I (T7EI) assay followed by Sanger sequencing for mutation detection. T7 endonuclease I assay with PCR products did not reveal the expected pattern of digestion for the insertion/ deletion (InDel) mutation. However, Sanger sequencing of the PCR products confirmed seven CRISPR/Cas9-based mutated T0 lines of LR-1 constructs and six CRISPR/Cas9-based mutated T0 lines of LR-2 constructs, with a percentage mutation frequency of 43.75 %, and 40 % for LR-1 and LR-2 constructs, respectively. The two LR-1 and LR-2 constructs targeting the RPK1 gene achieved similar mutation frequency. Overall, these results showed a high mutation efficiency of 41.93 %.
Vector
Target gene
Construct
sgRNA
No. of plants examined
No. of plants with mutations
% Mutation frequency
sgRNA GC content
CRISPR/ Cas9
TaRPK1
LR-1
sgRNA1 sgRNA2
16
07
43.75 %
41.67 %/ 45.83 %
LR-2
sgRNA1 sgRNA3
15
06
40 %
41.67 %/ 41.67 %
Total
31
13
41.93 %
3.4 Types of mutations caused by CRISPR/Cas9
The sequencing results showed two types of mutations, deletions, and insertions. The insertions of one bp, A, and T were detected at target sites (Figs. 3-6). The frequency of insertion was less as compared to that of deletion. Short deletions of one to nine bp were also observed. The deletions were mostly short, however longer deletions such as 12d, 17d, 19d, and 20d (Fig. 3A) were detected in T0-6, T0-10, T0-14, and T0-16 lines. The longer deletions were observed at target site 2 by sgRNA2 of LR-1 construct, showing high efficiency of sgRNA2 compared to sgRNA1 and sgRNA3 (Fig. 5A). The total deletion ratio was higher compared to total insertions and Cas9 cleavage sites usually 3 bp upstream of PAM when all the mutation events were summed. These findings indicate that our CRISPR/Cas9 system did work in the T0-edited plants.
CRISPR/Cas9 induced targeted mutagenesis of the TaRPK1 target gene by LR-1 construct within A, B, and D genomes of wheat T0 plants (A). PAM sites were shown in red, nucleotides in blue denote target site 1 (T1), and nucleotides in orange represent target site 2 (T2). The lowercase letter denotes insertion, dashes represent deletions and dots indicate nucleotides not shown. Spike architecture of wild type and edited transformed LR-1 T0 lines (B). Root architecture of wild type and edited transformed LR-1 T0 lines (C). (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
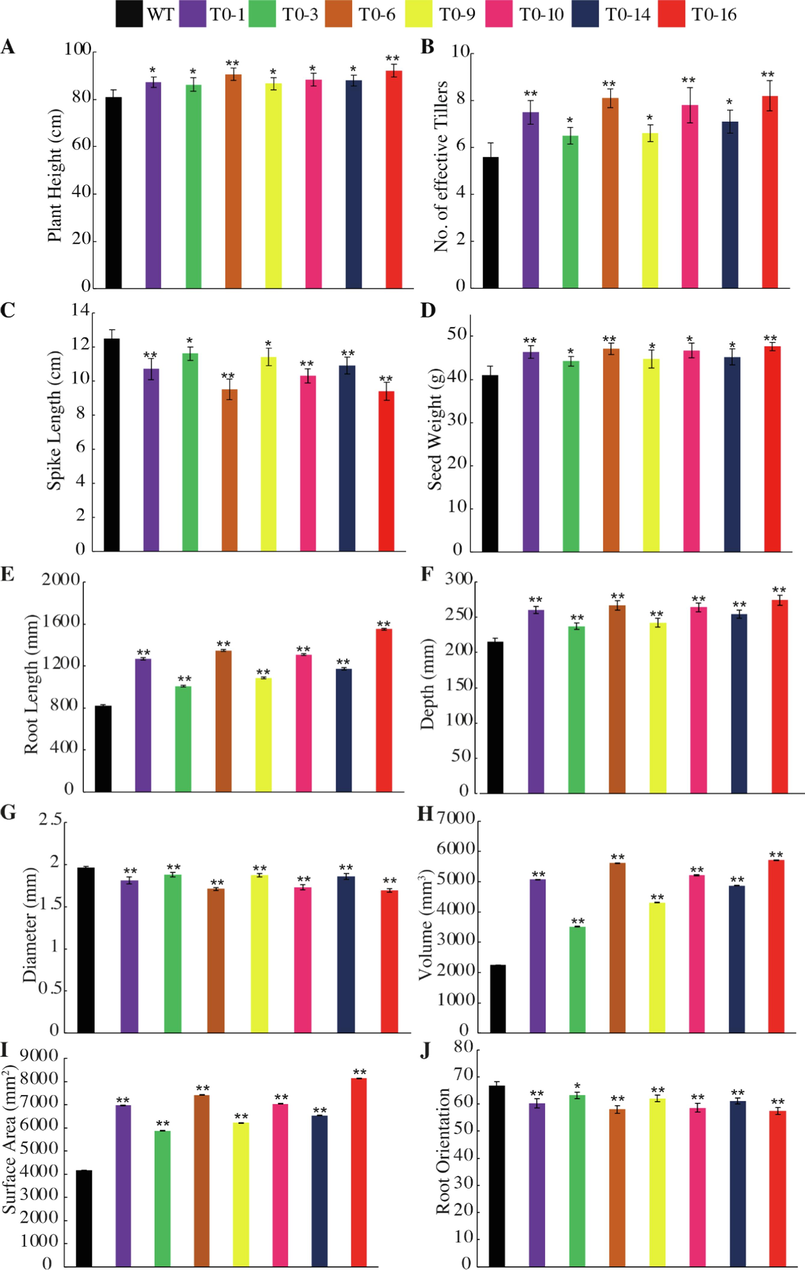
Phenotypic comparison of LR-1 construct for Plant height (cm, A), Number of effective tillers (B), Spike length (cm, C), Seed weight (g, D), Root length (mm, E), Root depth (mm, F), Root diameter (mm, G), Root volume (mm3, H), Root surface area (mm2, I), and Root orientation (J) of wild type (WT) and edited T0 lines. The data indicates the mean ± SE of three biological replicates. Student’s t-test was used (* = P < 0.05, and ** = P < 0.01).
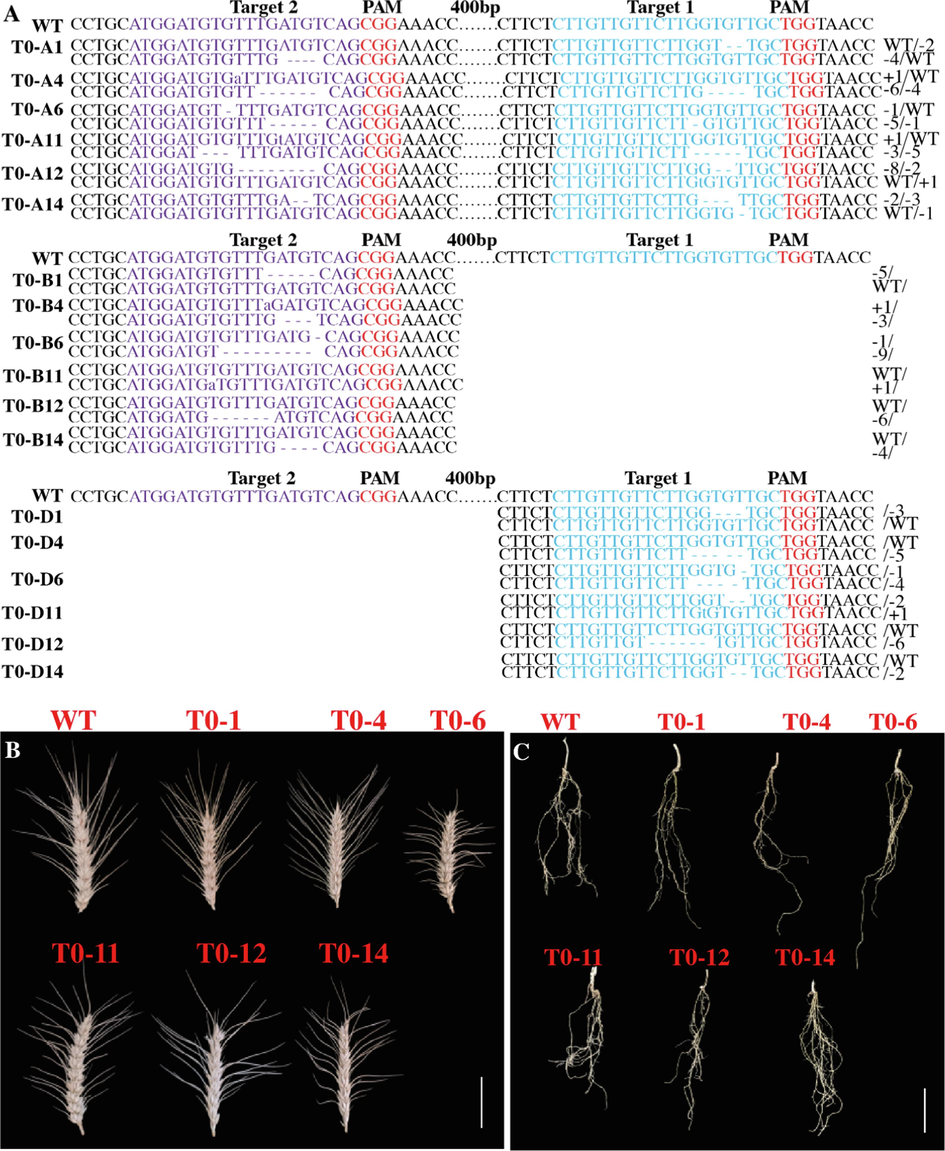
CRISPR/Cas9 induced targeted mutagenesis of the TaRPK1 target gene by LR-2 construct within A, B, and D genomes of wheat T0 plants (A). PAM sites were shown in red, nucleotides in blue denotes target site 1 (T1), and nucleotides in purple represent target site 2 (T2). The lowercase letter denote insertion, dashes represent deletions and dots indicates nucleotides not shown. Spike architecture of wild type and edited transformed LR-2 T0 lines (B). Root architecture of wild type and edited transformed LR-2 T0 lines (C). (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
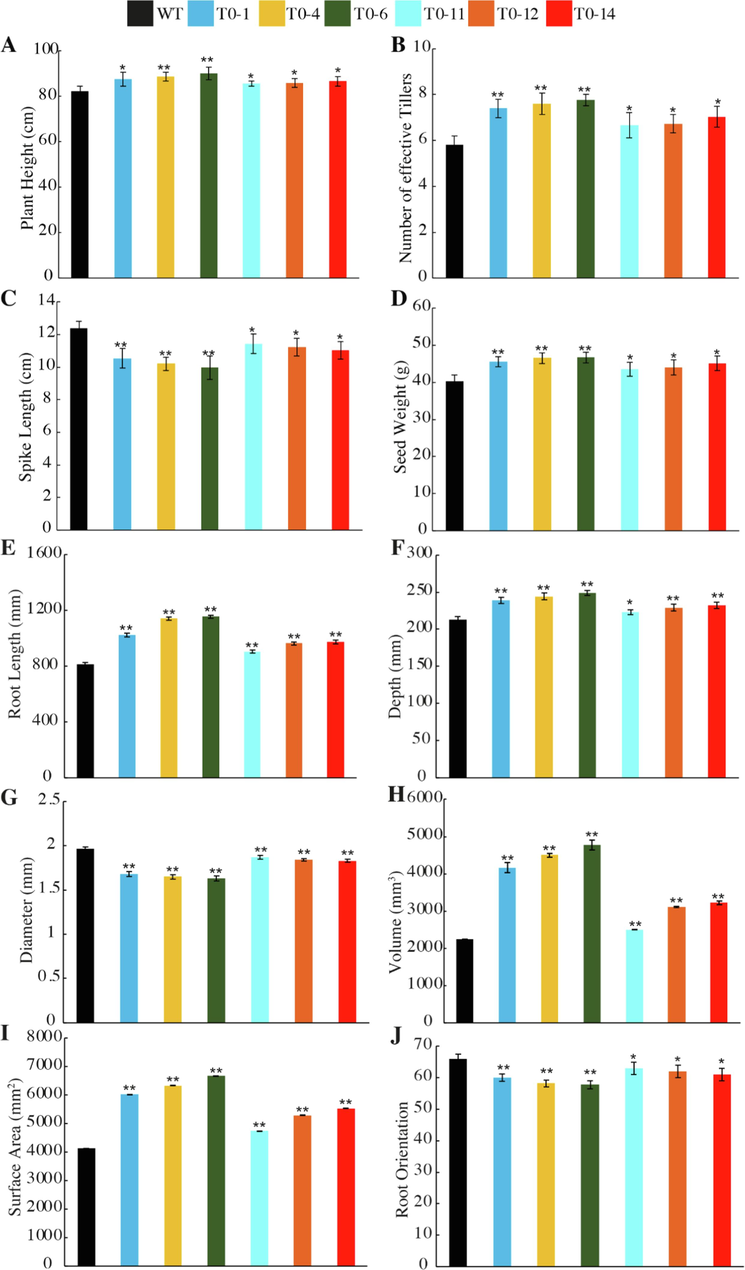
Phenotypic comparison of LR-2 construct for Plant height (cm, A), Number of effective tillers (B), Spike length (cm, C), Seed weight (g, D), Root length (mm, E), Root depth (mm, F), Root diameter (mm, G), Root volume (mm3, H), Root surface area (mm2, I), and Root orientation (J) of wild type (WT) and edited T0 lines. The data indicates the mean ± SE of three biological replicates. Student’s t-test was used (* = P < 0.05, and ** = P < 0.01).
3.5 Monoallelic and diallelic mutations in T0 generation
Given the complexity of possible results, from zero to three target sites and using 2 different constructs in complex sub-genomes of hexaploid bread wheat (Table 3). Results displayed higher monoallelic mutation compared to diallelic mutation. 37.5 % monoallelic mutation at target site 1 within the D genome by sgRNA1 was observed by the LR-1 construct (Table 3). The LR-2 constructs showed a higher monoallelic mutation frequency of 26.67 % at target sites 1 and 2 within the A, B, and D genomes (Table 3). The highest diallelic mutation of 25 % was observed at target site 2, by sgRNA2 of LR-1 construct within A genome of wheat. However, the 20 % higher diallelic mutation frequency was detected by sgRNA3 at target site 2 of the LR-2 construct within the A genome (Table 3).
Construct
Genome
sgRNA
No. of plants examined
Target sites on T0 plants
Genotype
Monoallelic mutation (MA)
Diallelic mutation (DA)
LR-1
A
sgRNA1
16
T 1
05 (31.25 %)
02 (12.5 %)
sgRNA2
T 2
03 (18.75 %)
04 (25 %)
B
sgRNA2
T 2
04 (25 %)
03 (18.75 %)
D
sgRNA1
T 1
06 (37.5 %)
01 (6.25)
sgRNA2
T 2
04 (25 %)
03 (18.75 %)
LR-2
A
sgRNA1
15
T 1
04 (26.67 %)
02 (13.33 %)
sgRNA3
T 2
03 (20 %)
03 (20 %)
B
sgRNA3
T 2
04 (26.67 %)
02 (13.33 %)
D
sgRNA1
T 1
04 (26.67 %)
02 (13.33 %)
3.6 Disruption of TaRPK1 effects morphological parameters and root system architectural traits
To see if TaRPK1 gene disruption effects the morphological parameters and root system architecture, we observed and measured these traits in edited lines along with control plants (Figs. 3-6). Alteration in spike architecture in the edited plants was observed (Fig. 3B). Plant height increased considerably in LR-1 edited T0 lines compared to WT plants (Fig. 4A). Plant height increased significantly in T0-16 and T0-6 lines. T0-16 had the most tillers number, followed by T0-6 > T0-10 > T0-1 LR-1 edited lines (Fig. 4B). Spike lengths were reduced significantly in genome-edited LR-1 lines (Fig. 4C). A significant reduction in spike length was observed in T0-16 lines. Data regarding 1000 grain weight was significantly increased in all of the T0-edited LR-1 lines with a maximum in T0-16 > T0-6 > T0-1 lines compared to the control plants (Fig. 4D).
We also detected LR-2 edited T0 wheat plants, which had changed spike architecture (Fig. 5B). The morphological traits such as the number of effective tillers and plant height were significantly increased, while the spike length was significantly reduced in LR-2 edited wheat T0-1, T0-4, and T0-6 plants (Fig. 6A-C). 1000 grain weight was significantly enhanced in all of T0 edited LR-2 wheat lines (Fig. 6D). All LR-2 T0 edited lines increased seed weight, with T0-1, T0-4, and T0-6 lines having the highest.
The root architectural traits showed significant effects in LR-1 genome-edited T0 lines (Fig. 3C). The root length, root depth, volume, and surface area were significantly increased (Fig. 4E-F and H-I), however, the root diameter and root angle were significantly reduced in genome-edited LR-1 (Fig. 4G and 4 J) T0 lines compared to the WT plants. T0-1, T0-6, T0-10, and T0-16 lines displayed very significant differences in comparison to the control plants. Similarly, in the other construct (LR-2), root architectural traits also displayed significant variations in genome-edited T0 lines (Fig. 5C).
Notably, editing of wheat TaRPK1 genes with specifically designed sgRNAs may have changed the open reading frame and amino acid sequence in transgenic edited lines relative to WT plants (Supplementary Figure S4). In comparison to WT plants, the reduced spike length, root diameter, and root angle and increased root length, depth, surface area, and volume increase the number of effective tillers and grain weight. Increased grain weight and effective tillers on modified lines offset the grain yield loss from lower spike length. This research shows that TaRPK1 editing improves root architecture and yield.
4 Discussion
Common wheat is an essential crop and has a complex genome comprising of A, B, and D sub-genomes (Zhang et al., 2018). Wheat genome editing is difficult and important for academics and scientists because of its intricacy (Zhang et al., 2019). The CRISPR/Cas9 system has advanced promptly recently (Ma and Liu, 2016). CRISPR/Cas9 was used to target wheat TaRPK1 genes (Fig. 1), from sgRNA design to agronomic characteristics analysis over T0 generation.
CRISPR/Cas9 has been used to generate double-strand breaks (DSBs). The principal repair mechanism is the Non-Homologous End Joining (NHEJ) mechanism. The frequency of NHEJ-mediated mutagenesis is influenced by a variety of factors. The major factors affecting the rate of this efficiency are the spatiotemporal expression of sgRNAs and Cas9 by promoters, selection of sgRNA, delivery methods of sgRNA /Cas9/ which leads to different numbers of transgenic copies, and, the sequence of a gene and its chromosomal position that affects the target gene accessibility of sgRNA/Cas9 complex to create DSBs (Horlbeck et al., 2016).
We adopted the two-sgRNAs-for-one-gene approach to enhance presumably double the rate of success or improve the probability of activation of at least one of the gRNA for mutagenesis, as reported in previous studies that not every synthesized sgRNA is highly active or active to create DSBs (Schmitz et al., 2020). This approach would enhance the attainment of targeted mutagenesis and larger deletion mutations. The gRNAs GC content is essential for the CRISPR/Cas9 efficiency (Zhang et al., 2018). In our study, the percentage of GC content in sgRNA1 and sgRNA3 was 41.67 %, and in gRNA2 was 45.83 % (Table 2). The gRNA2 having higher GC% content exhibited a higher efficiency (Zhang et al., 2018).
In the present study, we synthesized two constructs namely LR-1 and LR-2, with two gRNAs each, and successfully created mutagenesis in target sites of TaRPK1 genes, via Agrobacterium-mediated CRISPR/Cas9 in T0 plants. Our results indicated that the primary mutation type was deletion. Similar observations were also reported in other wheat studies (Ahmar and Gruszka, 2023). However, these observations contradict rice, where insertion is the dominant mutation type. We reported that the CRISPR/Cas9 system can achieve efficiently specific targeted editing with a mutation frequency of 43.75 % and 40 % for LR-1 and LR-2 constructs, respectively (Table 2). The stated efficiencies of mutations within different species or even different CRISPR platforms within wheat vary significantly. For instance, a relatively lower editing frequency of 5.6 %, to 20.8 %–54.2 % mutation efficiency was reported in T0 wheat plants (Zhang et al., 2019). 54.17 percent mutagenesis efficiency, far greater than previously reported, was found in bigger and more complex genomic cereal crops and wheat (Zhang et al., 2018). In contrast, a significant rate of editing up to 100 % have been reported by several laboratories working on rice (Viana et al., 2019). In dicotyledonous crops, such as soybean and tomato, CRISPR systems generated higher than 50 % mutation in T0 plants. In maize, 2 %–100 % editing efficiency was reported in the T0 plants (Zhu et al., 2016).
The yield is affected by numerous genes that influence abiotic stressors and root system architecture (RSA), including RPK1. Our findings support a prior rice research that found knockdown enhanced root system architecture, tillering counts, and plant height (Zou et al., 2014). In T0 wheat lines, CRISPR/Cas9 knockdown of the TaRPK1 gene altered root system architecture, spike architecture, tiller count, plant height, and grain weight. The LR-1 and LR-2 T0 lines have significantly altered morphological and architectural features relative to WT plants owing to CRISPR/Cas9-mediated genome editing insertions and deletions that altered the amino acid sequence and structure and function of TaRPK1. According to reports, root architecture and shoot system interactions help combat water shortage under drought stress and preserve production (Siddiqui et al., 2021). The mechanism of how RPK1 affects root system architecture might be via negative modulation of polar auxin transport (Fig. 7).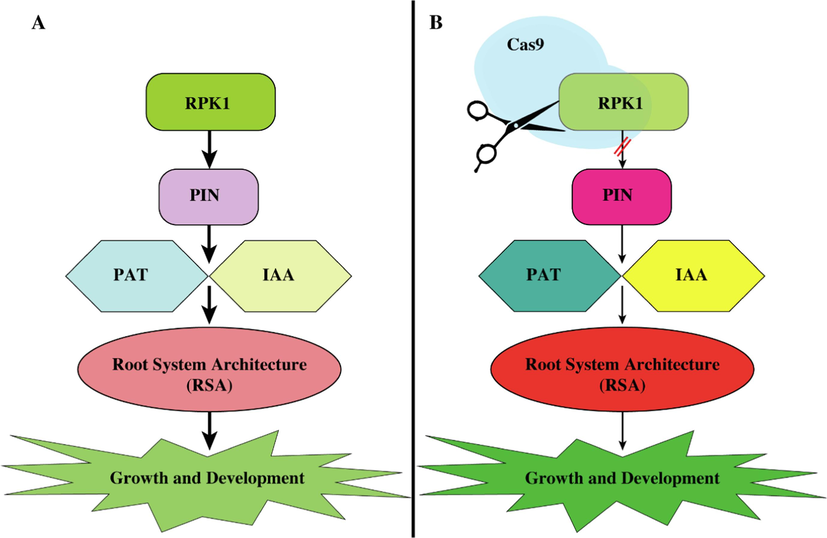
Involvement of RPK1 in root system architecture regulation. (A) normal RPK1 pathway, in which RPK1 is normally expressed and inhibits the expression of other genes such as PIN, IAA, and PAT. This less expression leads to poor root development. (B) Editing of RPK1 by using the Cas9 system leads to less expression of RPK1 and higher expression of PIN, PAT, and IAA. This causes the proper or better root development under drought stress.
5 Conclusion
In summary, the CRISPR/Cas9 system was highly efficient in generating TaRPK1 gene editing in wheat. TaRPK1 mutant lines with the necessary mutation were produced, and have different root architecture and agronomy. This research might boost bread wheat breeding. Our findings show that CRISPR's accuracy and speed will help enhance wheat at a pace that ensures global food security.
Funding
This research was conducted with the funds of the Agricultural Linkages Program of Pakistan (ALP # CS-399) and Sino-Pak Agricultural Breeding Innovation for Rapid Yield Enhancement (Project # 760). The funder has no role in designing or conducting any experiment or publication. Moreover, this work was supported by National Research Foundation of Korea (NRF) grant funded by the Korean Government (MSIT) (NRF-2021R1F1A1055482).
Acknowledgments
The authors extend their appreciation to Researchers Supporting Project number (RSP-2024 R369), King Saud University, Riyadh, Saudi Arabia.
Author contributions
AAR and MRK gestated the idea. AAR performed the research and prepared the first draft. MU, SF, KAA, and AMS visualized and updated the figures, improved the literature review, funding acquisition, and provided technical expertise to improve the article. AAR, NR, MU, and MRK helped in the write-up of the original draft. AAR, MU, SF, SHY and MRK helped in revision and funding acquisition. All authors carefully read, revised, and approved the article for submission.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Development of in planta transformation protocol for wheat. Afr. J. Biotechnol. 2011
- [Google Scholar]
- Genome-edited triple-recessive mutation alters seed dormancy in wheat. Cell Rep. 2019
- [Google Scholar]
- Crispr/cas9 boosts wheat yield by reducing brassinosteroid signaling. Trends Biochem. Sci 2023
- [Google Scholar]
- Crispr/cas9-mediated genome editing techniques and new breeding strategies in cereals–current status, improvements, and perspectives. Biotechnol. Adv. 2023
- [Google Scholar]
- Analyses of genes encoding the glycogen synthase kinases in rice and arabidopsis reveal mechanisms which regulate their expression during development and responses to abiotic stresses. Plant Sci. 2023
- [Google Scholar]
- A major root architecture qtl responding to water limitation in durum wheat. In: Frontiers in Plant Science. 2019.
- [Google Scholar]
- A streamlined protocol for wheat (triticum aestivum) protoplast isolation and transformation with crispr-cas ribonucleoprotein complexes. Frontiers in Plant Science; 2020.
- Char, S.N., Li, R., Yang, B., 2019. Crispr/cas9 for mutagenesis in rice. Transgenic plants, Springer: 279-293.
- Soil conditions affect cotton root distribution and cotton yield under mulched drip irrigation. Field Crop Res. 2020
- [Google Scholar]
- Calmodulin 1 regulates senescence and aba response in arabidopsis. In: Frontiers in Plant Science. 2018.
- [Google Scholar]
- Root length and root lipid composition contribute to drought tolerance of winter and spring wheat. Plant and Soil 2019
- [Google Scholar]
- Alien chromosome segment from aegilops speltoides and dasypyrum villosum increases drought tolerance in wheat via profuse and deep root system. BMC Plant Biol. 2019
- [Google Scholar]
- Effects of breeding history and crop management on the root architecture of wheat. Plant and Soil 2020
- [Google Scholar]
- Dosage differences in 12-oxophytodienoate reductase genes modulate wheat root growth. Nat. Commun. 2023
- [Google Scholar]
- Horlbeck, M.A., Witkowsky, L.B., Guglielmi, B., Replogle, J.M., Gilbert, L.A., Villalta, J.E., Torigoe, S.E., Tjian, R., Weissman, J.S., 2016. Nucleosomes impede cas9 access to DNA in vivo and in vitro. elife.
- Kim, D., Alptekin, B., Budak, H., 2018. Crispr/cas9 genome editing in wheat. Functional & integrative genomics.
- Crispr/cas9-mediated atgata25 mutant represents a novel model for regulating hypocotyl elongation in arabidopsis thaliana. Mol. Biol. Rep. 2023
- [Google Scholar]
- Crispr/cas9-based genome editing and its applications for functional genomic analyses in plants. Small Methods 2019
- [Google Scholar]
- Generation of paternal haploids in wheat by genome editing of the centromeric histone cenh3. Nat. Biotechnol. 2020
- [Google Scholar]
- Ma, X., Liu, Y.G., 2016. Crispr/cas9‐based multiplex genome editing in monocot and dicot plants. Current protocols in molecular biology.
- Application of crispr-cas12a temperature sensitivity for improved genome editing in rice, maize, and arabidopsis. BMC Biol. 2019
- [CrossRef] [Google Scholar]
- A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant 1962
- [Google Scholar]
- Root and canopy traits and adaptability genes explain drought tolerance responses in winter wheat. PLoS One 2021
- [CrossRef] [Google Scholar]
- Wheat root systems as a breeding target for climate resilience. Theor. Appl. Genet. 2021
- [Google Scholar]
- Genome-wide identification and characterization of receptor-like protein kinase 1 (rpk1) gene family in triticum aestivum under drought stress. Front. Genet. 2022
- [Google Scholar]
- Exploring the genetic architecture of root-related traits in mediterranean bread wheat landraces by genome-wide association analysis. Agronomy 2020
- [Google Scholar]
- Genome-wide analysis of mikc-type mads-box genes in wheat: Pervasive duplications, functional conservation and putative neofunctionalization. New Phytol. 2020
- [Google Scholar]
- Schmitz, D.J., Ali, Z., Wang, C., Aljedaani, F., Hooykaas, P.J., Mahfouz, M., de Pater, S., 2020. Crispr/cas9 mutagenesis by translocation of cas9 protein into plant cells via the agrobacterium type iv secretion system. Frontiers in genome editing.
- Rhizovision explorer: Open-source software for root image analysis and measurement standardization. AoB plants; 2021.
- Overexpression of the receptor-like protein kinase genes atrpk1 and osrpk1 reduces the salt tolerance of arabidopsis thaliana. Plant Sci. 2014
- [Google Scholar]
- Genetics and genomics of root system variation in adaptation to drought stress in cereal crops. J. Exp. Bot. 2021
- [Google Scholar]
- Narrow leaf21, encoding ribosomal protein rps3a, controls leaf development in rice. Plant Physiol. 2021
- [Google Scholar]
- Mutagenesis in rice: The basis for breeding a new super plant. Frontiers in Plant Science; 2019.
- Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 2014
- [Google Scholar]
- Zhang, S., Zhang, R., Gao, J., Gu, T., Song, G., Li, W., Li, D., Li, Y., Li, G., 2019. Highly efficient and heritable targeted mutagenesis in wheat via the agrobacterium tumefaciens-mediated crispr/cas9 system. International journal of molecular sciences.
- Analysis of the functions of ta gw 2 homoeologs in wheat grain weight and protein content traits. Plant J. 2018
- [Google Scholar]
- Targeted mutagenesis using the agrobacterium tumefaciens-mediated crispr-cas9 system in common wheat. BMC Plant Biol. 2018
- [Google Scholar]
- Efficiency and inheritance of targeted mutagenesis in maize using crispr-cas9. J. Genet. Genomics 2016
- [Google Scholar]
- Zou, Y., Liu, X., Wang, Q., Chen, Y., Liu, C., Qiu, Y., Zhang, W., 2014. Osrpk1, a novel leucine-rich repeat receptor-like kinase, negatively regulates polar auxin transport and root development in rice. Biochimica et Biophysica Acta (BBA)-General Subjects.
Appendix A
Supplementary material
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jksus.2023.103063.
Appendix A
Supplementary material
The following are the Supplementary data to this article:
Supplementary data 1
Supplementary data 1
S1: A flowchart of targeted mutagenesis in bread wheat. S2: Construction of gRNA expression cassettes. S3: Genomic DNA of control (1) and transgenic plants (2-4). The marker was denoted with “M”. S4: Multiple sequence alignment of TaRPK1 proteins depicting structural predictions
Supplementary data 2
Supplementary data 2
Supplementary data 3
Supplementary data 3
Supplementary data 4
Supplementary data 4
S1: The RPK1 sequences in the A, B, and D sub-genomes of wheat. S2. Alignment of A, B, and D CDS genomes. S3: Primer sequences used for sequencing of A, B, and D genomes







