Translate this page into:
Morphological, physico-biochemical and marker-based diversity of desi cotton (Gossypium herbaceum L.) germplasm
⁎Corresponding author. sushil254386@yahoo.com (Sushil Kumar)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
Desi cotton (Gossypium herbaceum) is one of the important crops as it is valuable source of lint and spinnable fibre for textile industries. G. herbaceum is resistant to biotic and abiotic stresses and is sturdy crops species. The genetic diversity analysis of this crop is important for further improvement in its productivity to cap the gap between demand and supply of short staple cotton. Therefore, a set of 48 inbred lines of desi cotton were used in this study and their variability was estimated using morphological traits, yield parameters, seed physical properties, fibre quality parameters and seed chemical parameters. Similarly, 13 SSR and five ISSR markers were used for molecular diversity evaluation of germplasm. In this study, ANOVA showed significant differences among all genotypes for all the traits, demonstrating a substantial amount of genetic variability in studied genotypes. Morphological studies showed that genotype Radhanpur had higher seed yield (185.50 g/plant) and lint yield (68.60 g/plant). Physico-biochemical studies suggest that genotypes GVhv-845 had higher fibre strength (26.10 g/tex), Wagad (19.05%) was higher in oil and W8 (51.67%) had maximum seed protein. The Manhattan dissimilarity co-efficient based phenotypic diversity generated six main clusters. The average dissimilarity value among genotypes was 0.30, indicating moderate phenotypic variability. The dendrogram generated from pooled data of SSR and ISSR markers based on Jaccard’s similarity matrix grouped the genotypes into four main clusters. The genetic coefficient of similarity among the genotypes ranged from 0.15 to 0.70 with an average of 0.32. The present study revealed a low correlation between phenotypic and marker-based matrices(r = 0.09). Along with, low correlation, both the matrices placed a limited number of genotypes (nine) in the same clusters in their respective dendrograms. The low correlation indicated that the two methods were different and highly variable as molecular markers are neutral in behave than quantitative traits.
Keywords
ANOVA
Diploid cotton
SSR
ISSR
Fibre strength
Oil
1 Introduction
Cotton, renowned as “white gold”, is a well-known fibre crop. It belongs to the genus Gossypium and family Malvaceae. Desi or levant cotton (G. herbaceum) has a 2n = 26 chromosomes and belongs to "A" genome (Fryxell, 1979; Fryxell, 1992). G. herbaceum has three varieties namely typicum, fruitescens and africanum, and five geographic races viz. persicum, kuljianum, acerifolium, wightianum and africanum of which wightianum was grown in western India known for drought resistance.
Presently, tetraploid cotton especially American cotton covers > 90% area of world cotton due to its superior fibre quality and productivity; and has achieved the status of primary cotton. Diploid cottons are cultivated solitary in traditional cotton-cultivating regions of Asian countries like India, Pakistan, China, Bangladesh and Iran (Kulkarni et al., 2009). Levant (G. herbaceum) and tree (G. arboreum) cotton together provide two percent of the global cotton (Zhang et al., 2008). G. herbaceum is tolerant to various abiotic stresses like salinity, drought and wind (Parekh et al., 2018). India occupies 37.56% of the world cotton area and produces 24.26% of world cotton production (Prakash, 2020). This species is suitable for rainfed areas but fewer undesirable traits like low fibre quality, small boll size, undesirable plant frame makes it less suitable for cultivation (Parekh et al. 2016). Therefore, there is a need for an extensive evaluation of germplasm and adaptive performance of the species.
Recently interest in cottonseed has extended as feed for non-ruminants or even human consumption, because cotton seeds are rich in protein (42% of dry matter) and oil (20% of dry matter). Cottonseed meal is rich in amino acid composition i.e., lysine (1.70%-2.00%), tryptophan (0.51%-0.53%), methionine (0.70%-1.00%) and also rich in carbohydrates. Across the globe, 50 to 55 million tons of cotton-seed are produced every year with a value of 7 to 8 billion US dollars (Narayanan et al., 2014). Cotton is an important oilseed in the world which is genetically modified for oil extraction next to soybean, corn and rapeseed. It is also cellulose source for linters for manufacturing numerous personal care goods and high-quality paper for currency printing (Wikipedia 2013 Euro bank Notepaper and currency economy in the world). Gossypol, a lipid-soluble secondary phytochemical, is a naturally occurring polyphenol confined in the lysigenous glands beneath the palisade cells of the leaves and hypodermal cells of stem and bolls in all the species (Dowd, 2003). Gossypol, a secondary metabolite, was first discovered by J. J. Longmore in 1886 and purified in crystal form by L. Marchlewski in 1889 (Croteau et al., 2000). It is important phytochemical component of interest due to its various properties such as anti-cancer, anti-microbial, anti-oxidant and male contraceptive (Anonymous, 1995). Glanded cotton generally contains gossypol while glandless cotton has low or no gossypol content (Punit et al., 1991). Many breeding efforts were initiated for the production of cotton seed varieties with low gossypol content.
With a projected nine billion human population, global textile fibre requirements are projected at 180 million metric tons. The share of cotton should be increased to 40 to 50 million metric tons by 2050 (Narayanan et al., 2014). Cotton lints are short fibres left after removal of long fibres; these are yet another important by-product as they are a source of purest cellulose. Usually, desi cotton contains 4.30% to 5.90% of cellulose. Hull, the seed coat, constitutes about 37% of the seed weight.
For any crop improvement programme, analysis of genetic diversity is prerequisite. To have a reliable estimate of genetic relationships and genetic diversity polymorphic (morphological or DNA) markers are required. Most of genetic diversity analysis studies in desi cotton have been carried out using morphological markers only. Now-a-days Polymerase Chain Reaction (PCR)-based molecular markers especially simple sequence repeats (SSR) and inter simple sequence repeats (ISSR) are time-tested powerful tools to analyse genetic relationships and genetic diversity. Considering the importance of cotton as an industrial crop, the current study was formulated to assess the genetic variability among the genotypes of G. herbaceum.
2 Material and methods
2.1 Experimental materials and field evaluation
The experimental site was located in Regional Cotton Research Station, Viramgam. It has semi-arid climate with an annual average rainfall of 715 mm distributed and has medium black soil. Seeds of the forty-eight inbred lines (Suppl. Table 1) were planted in randomized complete block design (RBD) in two replications during kharif 2020–21, following 120 × 30 cm spacing. The package of practices was followed as per the recommendations for raising the healthy crop.
2.2 Morphological characterization
The data on eight morphological characters namely 50% Flowering (DFF, days), plant height (PH, cm), bolls/plant, five bolls weight (g), seed index (g), seed cotton yield/plant (g), lint yield/plant (g), and ginning (%) were recorded. The phenotyping for studied traits were carried out on five competitive plants per genotype in each replication. Phenotypic data DFF and days to maturity (DM) were collected on a plot basis.
2.3 Physico-biochemical characterization
2.3.1 Physical characterization of seed
Seeds were acid de-linted using concentrated H2SO4 as per Karivaradaraaju (2007). Fluid displacement method was used to measure average unit volume (mm3) of seeds (Perea-Flores et al., 2011). Bulk density (gcm−3), true density (gcm−3, toluene displacement method) and porosity (%) of the seeds was determined as suggested by Mohsenin (1970).
2.3.2 Measurement of fibre quality traits
Fibre samples (100gm) were sent to Central Institute for Research on Cotton Technology (CIRCOT) lab, Navsari Agricultural University (NAU), Surat for quality measurement. The pooled fibre sample was analyzed for fibre quality traits. The sample testing was carried out at 65 ± 2% relative humidity and 27 ± 2 °C temperature in HVI (High Volume Instrument) mode for the upper-half mean fibre length (UHML, mm), fibre strength (g/tex), fibre fineness (µg/inch), fibre uniformity (FU, %) and fibre elongation (FE, %).
2.3.3 Seed oil and crude protein
Total oil extracted with a Soxhlet apparatus using hexane (AOAC, 1965). For oil measurement, 0.5g of ground cotton seed powder (delinted) was used. Micro-Kjeldahl method was deployed to estimate crude protein (AOAC, 1965). A total of 40 mg defatted sample (cotton seed powder, collected after oil extraction) was used for protein measurement.
2.3.4 Gossypol content (%)
Seed gossypol was determined by Reverse Phase Liquid Chromatography- Mass Spectrometry (RPLC-MS). Gossypol standard (G8761) was procured from Sigma – Aldrich. Complex reagent required for gossypol estimation was prepared where 2 mL of 3-aminopropanol and 10 mL of glacial acetic acid were mixed in a beaker and final volume made up to 100 mL using N, N-dimethyl formamide. Then, one gram of fine cotton seed powder was mixed with 25 mL of complex reagent and kept in hot water bath at 100 °C for 2 h. The yellow colour developed in the complex reagent indicates the presence of gossypol. Then cocktail was filtered using 0.2 mm filter and was diluted with methanol (1:4). At that point 100 µL diluted sample was further diluted with 900 µL of methanol and used for RPLC-MS on EkspertTM Ultra LC 100-XL system (Eksigent, USA) (Karishma et al., 2016). Detailed methodology of RPLC-MS and representative chromatogram is given in supplementary document 1.
2.4 DNA extraction and marker amplification
The genomic DNA from freshly collected bulked leaf tissues was extracted using the CTAB (Cetyl Trimethyl Ammonium Bromide) method, as previously described by Doyle and Doyle (1990). The quantity of good quality DNA was quantified in Nanodrop ND-1000 (Thermo, USA). DNA samples were diluted to a concentration of 30 ng µL−1 with TE. SSR and ISSR amplification in thermal cycler (SensoQuest, Germany) was achieved as per Rukhsar et al. (2017) and Patel et al. (2015), respectively, in 10 mL PCR mix having 1.5 mL DNA, 5 mL Master Mix (2×) (Takara, Japan), and 1 mL of 10pMol primer and 2.5 mL nuclease free water. The PCR products were separated on 1.5% agarose gel and 6% non-denaturing polyacrylamide gel electrophoresis (PAGE) followed by silver staining (Goldman and Merril, 1982). Stained gels were documented through scanner (Bio-6000, Microtek, Taiwan) at 600 dpi. The size of amplified band was estimated by matching it with a 100 bp ladder.
2.5 Statistical analysis of data
The data recorded for studied traits were used to analyse the variance (ANOVA) and critical difference (CD) to distinguish significantly dissimilar genotypes. Though, SSR is co-dominant, but for pooled analysis of both marker systems, we scored SSR data like a dominant marker (0/1 fashion) and consider each amplicon as an autonomous locus. The bands of both marker systems were recorded as 1 (band present) or 0 (band absent). This data was used to compute the polymorphism information content (PIC) and resolving power (Rp) value as suggested by Sharma et al. (2017). Multiplex ratio, effective multiplex ratio and marker index were calculated as per Powell et al. (1996). Jaccard’s similarity (J) coefficient was calculated to describe the genetic similarity among genotypes and the UPGMA dendrogram was prepared using SAHN algorithm with NTSYSpc (Rohlf, 2005).
3 Result and discussion
3.1 Analysis of variance (ANOVA)
Genetic variability is the total of genetic differences found among the individuals of a population. A diversity study is a quantitative estimation of genetic differences. Table 1 represents the analysis of variance for morphological and physico-biochemical characters indicating the mean sum of squares. In this study, genotypic differences were highly significant for all the characters, indicating a considerable amount of genetic variability among the studied genotypes. The presence of variability among these genotypes suggests ample scope for the selection of superior genotypes which can be used directly as a variety or as parents in plant breeding programmes. *and ** indicate significant at 5% and 1% levels of probability, respectively. Fuzzy seeds genotype: GVhv 1057.
Trait
Source of variation and Mean squares
Mean Performance
Replication (df = 1)
Genotypes (df = 47)
Error (df = 47)
Mean
Range
±S. Em
CD (0.05)
CV%
Days to 50% flowering
4.17*
59.33**
0.64
97.38
83.50 (9721)-107.50 (Radhanpur)
0.40
1.6
0.82
Plant height (cm)
137.76
885.26**
132.16
143.81
97.70 (Wagad 8)-199.30 (9759)
5.70
23.12
7.99
Bolls per plant
884.52*
151.33**
68.52
38.61
19.70 (BKhv 8)-62.00 (Radhanpur)
4.14
16.65
21.44
Five bolls weight (g)
0.86
5.44**
1.27
10.84
4.45 (Fuzzy seeds) −14.20 (9761)
0.56
2.26
10.38
Seed index (g)
0.02
0.62**
0.31
6.1
4.75 (Fuzzy seeds) −7.25 (9762)
0.28
1.11
9.09
Seed cotton yield per plant (g)
7086.40*
1797.50**
689.4
87.96
31.40 (Red kalyan) −185.50 (Radhanpur)
13.10
52.82
29.85
Lint yield per plant (g)
872.42*
249.40**
82.82
31.7
0.00 (Fuzzy seeds) − 68.60 (Radhanpur)
4.55
18.30
28.71
Ginning percentage
2.59
88.27**
5.52
36.19
0.00 (Fuzzy seeds) −45.15 (K 16)
1.18
4.73
6.49
Average unit volume(mm3)
0.91*
5.64**
0.18
5.65
3.33 (Gheti)-9.17
0.30
0.55
7.50
Bulk density (gcm−3)
2220.78*
2947.74**
492.52
0.469
0.32 (KS 5) −0.53 (9721)
15.69
44.64
4.73
True density (gcm−3)
185.2
11140.00*
2550.1
0.986
0.86 (Wagotar) −1.24 (Dhumad)
55.51
157.91
7.96
Porosity (%)
1.05
49.64**
13.86
52.09
43.99 (Wagotar) −68.04 (KS 5)
2.63
7.49
7.15
Upper half mean length (mm)
0.03
47.47**
0.12
21.93
0.00 (Fuzzy seeds) −25.15 (GVhv 149)
0.24
0.69
1.58
Fibre strength (g/tex)
0.59
45.49**
0.28
20.6
0.00 (Fuzzy seeds) −26.10 (GVhv 845)
0.38
1.06
2.57
Uniformity index (%)
0.09
531.83**
0.37
77.11
0.00 (Fuzzy seeds) −83.50 (Bhv 25)
0.05
0.15
1.55
Fibre fineness (µg/inch)
0.0001
2.59**
0.0056
4.86
0.00 (Fuzzy seeds) −7.00 (9757)
0.43
1.22
0.79
Elongation (%)
0.0001
2.39**
0.0052
5.15
0.00 (Fuzzy seeds) −5.65 (GVhv 845)
0.05
0.15
1.40
Oil content (%)
23.08*
28.69**
28.69
21.69
10.02 (1502)-19.05 (Wagad)
2.47
7.03
16.11
Protein content (%)
19.88
68.55**
6.82
41.78
19.18 (9761)-51.67 (W 8)
5.52
14.91
18.7
Gossypol content (%)
0.06*
0.29**
0.01
0.78
0.33 (Fuzzy seeds)-2.39 (GVhv 149)
0.08
0.23
14.62
3.2 Character variance analysis
3.2.1 Morphological parameters
Days to 50% flowering, an indicator of the earliness, is an important trait in desi cotton as it is mainly cultivated in rainfed araes. In the current study, DFF ranged from 83.50 (9721) to 106.50 (Radhanpur) days with a mean of 97.38 days. A similar mean value was observed by Patel et al. (2016) in G. herbaceum (95.10 days). Dwarfness, opposite to tall plant height, is desirable trait as dwarf plants are suitable for high-density planting, thus increasing the yield. The plant height (PH) in current study ranged from 97.70 cm (Wagad-8) to 199.30 cm (9759) with an average of 144.00 cm. In earlier reports, different plant height in cotton germplasm was observed like 166 cm by Patel et al. (2016) and 94.80 cm by Gapare et al. (2017). Bolls per plant are directly related to higher seed cotton yield (Ahmad et al., 2011). The average bolls per plant were 38.61. Wagad-8, a dwarf genotype, demonstrated a minimum number of bolls (19.70 bolls). The average number of bolls per plant was higher than earlier report (34.14 by Ranjan et al., 2014). Lower number of bolls per plant was also earlier recorded by Erande et al. (2014) in G. arboreum (14.76 bolls). Boll weight is directly associated with yield hence higher boll weight is desirable trait. In the present study, weight of five bolls ranged from 4.45 g (GVhv 1057: a Fuzzy seeds genotype) to 14.20 g (9761). A similar range and mean were obtained by Dhivya et al. (2014; 4.30 to 12.40 g).
3.2.2 Yield parameters
Seed index is important in determining yield and oil content. The average seed index of the forty-eight genotype was found to be 6.09 g, ranging from 4.75 g (Fuzzy seeds) to 7.00 g (9762). These results are in agreement with Erande et al. (2014; 6.93 g) and Saravanan et al. (2021; 5.29 g). Yield, a complex trait, is influenced by many genes and modified by environmental conditions. The seed yield was between 31.40 g (Red Kalyan) and 185.50 g (Radhanpur) with a mean of 87.96 g per plant suggesting a considerable variation in germplasm. In this study, lint yield per plant ranged from zero (Fuzzy seeds) to 68.60 g (Radhanpur). Similar range and mean were noticed by Snider et al. (2016; 1171 to 1534 Kg ha−1) and Zeng and Meredith (2009; 769 Kg ha−1). Contrastingly, Erande et al. (2014; 10.90 g) observed lower lint yield. Genetic studies indicated ginning trait is governed by additive gene action. Ginning percentage was 36.19%. A similar mean was recorded by Gapare et al. (2017; 39.70%) and Dhivya et al. (2014; 37.30%). A slightly higher ginning percentage was recorded by Patel et al. (2016; 45.60%).
3.2.3 Seed physical parameters
The average unit volume of observed genotypes ranged from 3.33 to 9.17 with an average of 5.65 mm3. The true and bulk densities are considered for designing seed hopper dimensions in ordinary seed planters (Ramesh et al., 2015). The true density of the observed genotypes ranged from 0.86 to 1.24 (Dhumad) with an average of 0.99gcm−3. Though, Ramesh et al. (2015; 1.11 gcm−3) have reported similar observations. In the current study, the bulk density ranged between 0.32 (KS-5) and 0.53 (9721) with a mean value of 0.47 gcm−3. Earlier, Vinayaka et al. (2020; range 0.545 to 0.613 gcm−3), and Ramesh et al. (2015; 0.589 gcm−3) estimated similar mean bulk density. The average value of porosity was observed to be 52.09% and the value ranged from 43.98 (Wagotar) to 68.04% (KS-15). The genotypes with low porosity will dry very slowly. Prior to this study, Ramesh et al. (2015) have reported a similar mean (46.76%) for porosity.
3.2.4 Fibre quality parameters
UHML (upper half mean length) below 24 mm is not considered good quality fibre. The mean value of the UHML was 21.93 mm ranging between zero (Fuzzy seeds) and 25.15 mm (GVhv-149). Genotypes namely 4651/3189 (25.05), GVhv 845 (25.05), WJ 80 (24.85), Bkhv-9 (24.75) and Blach-1 (24.70) were statistically at par with GVhv149. However, earlier Wang et al. (2013) recorded a higher mean value of 35.35 mm. For good spinning in modern high-throughput machineries, high tensile strength in fibres is required. The fibres of GVhv-845 (26.10 g/tex) has maximum tensile strength. Uniformity (U) is a crucial fibre quality characteristic determining the maturity of fibres. Higher U value indicates that the thread spun from such fibres is constant in size and strength, with small waste of fibres and is affected by the environment. U was maximum in Bhv-25 (83.50) while mean was 77.11% and it was similar to earlier reports (Gapare et al. 2017; Wang et al. 2013). Micronaire measures the fineness of fibre and lower value is desired. It governs the texture of fibre into soft/coarse and silky/harsh and is affected by the environment. Micronaire was acceptable with a mean of 4.86 µg/inch. Similar results were noticed by Dhivya et al. (2014; 4.30 µg/inch).
3.2.5 Seed chemical parameters
With high level of tocopherol, cotton is the sixth primary source of edible oil source around the world (Shahrajabian et al., 2020). The oil content ranged between 10.15% (1502) to 29.05% (Wagad). A similar range was shown by Kulkarni et al. (2009; 20.4%). A renewed interest of cotton seed oils simultaneously led to a cotton seed cake by-product that is left after oil extraction. This can be fed to livestock as it contains higher amount of protein (G. arbeoreum = 34.00%, G. herbaceum = 34.50%, G.hirsutum = 36.20% and G. barbadense = 36.20%). The average value of protein content among the genotypes was 41.24%, ranging between 19.18% (9761) and 51.67% (W8). The lower content of gossypol is desirable for edible oil. The average gossypol content of the forty-eight genotypes was found to be 0.79%, ranging from 0.33% (Fuzzy seeds) to 2.37% (GVhv 149). Pandey and Thejappa (1975) carried out gossypol estimation following the norms of AOCS (American Oil Chemists’ Society) and reported 1.36% of average gossypol value from different species under study. Scheffler and Romano (2008) recorded an average value of 1.31% of gossypol content using dry seeds of upland cotton.
3.3 Phenotypic diversity analysis
The clustering based on Manhattan dissimilarity coefficient divided the genotypes into six main clusters at a cut-off value of 0.30 (Fig. 1, Table 2) indicating moderate phenotypic variability. The dissimilarity between genotypes ranged from 0.11 (4651/3189 and 509–4-White) to 0.99 (4651/3189 and 615) indicating the highest and lowest similarity for the respective pair of genotypes.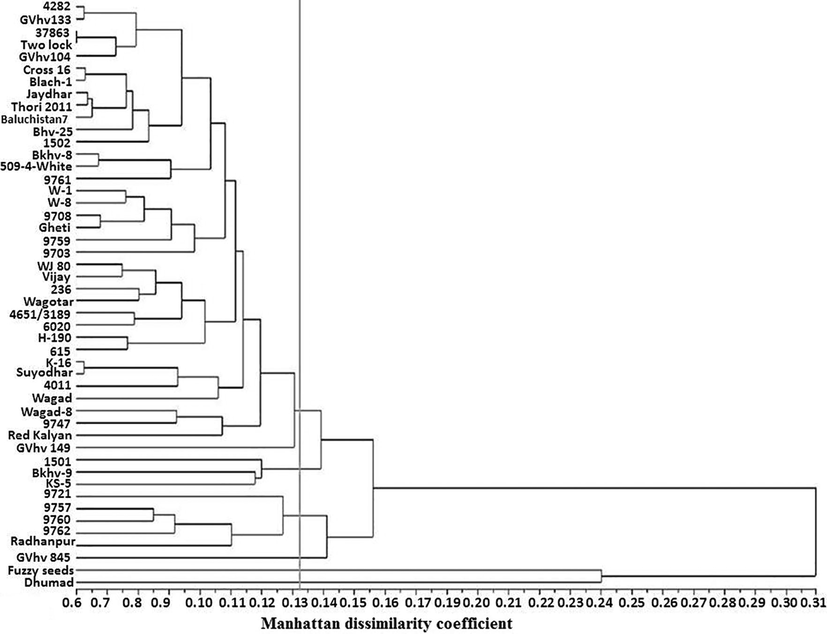
Grouping of 48 cotton genotypes based on 20 morphological traits using Manhattan distance.
Cluster No.
I
II
III
IV
V
VI
Genotype(s) per cluster
37.0
3.0
5.0
1.0
(GVhv-845)1.0
(GVhv 1057)1.0
(Dhumad)
Days to 50% flowering
96.82
100.33
99.50
105.50
100.50
87.00
Plant height (cm)
139.74
146.13
172.14
150.60
135.00
148.10
Bolls per plant
35.82
48.90
52.58
44.00
38.90
35.60
Five bolls weight (g)
11.06
11.47
10.40
7.75
4.45
12.40
Seed index (g)
6.13
6.35
6.00
6.40
4.75
5.70
Seed cotton yield per plant (g)
77.87
101.77
157.02
116.50
50.80
83.40
Lint yield per plant (g)
28.78
42.60
52.76
36.70
0.00
28.70
Ginning percentage
37.25
41.61
33.68
31.51
0.00
34.43
Average unit volume (mm3)
5.73
5.00
5.65
3.92
6.67
5.17
Bulk density (gcm−3)
471.52
400.21
498.37
516.67
397.74
473.78
True density (gcm−3)
965.66
976.80
1053.28
1020.00
900.00
1239.79
Porosity (%)
51.18
60.19
52.63
49.25
56.69
57.10
Upper half mean length (mm)
23.07
23.65
20.59
25.05
0.00
21.013
Fibre strength (g/tex)
21.52
22.18
19.98
26.10
0.00
22.10
Fibre fineness (µg/inch)
4.80
4.00
6.43
4.90
0.00
6.40
Uniformity index (%)
80.57
80.67
79.30
82.00
0.00
74.90
Elongation (%)
5.38
5.40
5.32
5.65
0.00
–
Oil content (%)
17.85
15.51
18.81
18.59
18.72
19.98
Protein content (%)
41.58
39.38
42.89
31.08
37.64
39.83
Gossypol content (%)
0.46
1.47
0.21
0.14
0.13
0.18
Cluster I comprised thirty-seven genotypes, indicating a moderate level of variation in all the parameters under study. Cluster II consists of three genotypes (1501, KS-5 and BKhv-9), characterized by high values of boll number, boll weight, seed index, lint yield and gossypol content, low values of bulk density and oil content. Cluster III contains five genotypes, namely 9721, 9757, 9760, 9762 and Radhanpur. This cluster is characterised by more boll number, good seed and lint yield, tall plants, higher fibre fineness and oil content. Cluster IV has only one genotype (GVhv 845) which has mean values for characters like days to 50% flowering, boll number, seed index, seed yield, true density, bulk density, UHML, fibre strength and oil content. Further the cluster V has one genotype (Fuzzy seed) which is characterized by low boll weight, seed index, seed yield, bulk and true density, oil and gossypol content and did not yielded lint. Lastly the cluster VI consists of only one genotype (Dhumad) which is an early maturing variety with low seed index, oil and gossypol content and also characterized by short fibres. Pundir (2019) also reported similar pattern of diversity where sixty genotypes of G. arboretum were divided into seven clusters based multivariate analysis (Mahalanobis’s D2 statistics). Dhanalakshmi (2011) also reported that Mahalanobis’s D2 statistics-based divergence analysis clustered the fifty-seven desi cotton genotypes into eight clusters.
3.4 Genotypic diversity analysis
3.4.1 SSR marker
A set of 49 SSR primers were screened on two DNA samples to identify polymorphic primers. These primer pairs were selected for the present study following the works of Parekh et al. (2016), Yu et al. (2008) and Sethi et al. (2016). Out of 49, 32 (65.3%) primers showed amplification. 13 (40.62%) primers out of 32 proved to be polymorphic (Table 3). The 13 primers generated 70 alleles with band size ranging from 94 (CES-21 and CES-28) to 432 bp (CES-8). Similarly, Mishra et al. (2013) reported alleles with band size 80–655 bp, Ezhilkumar and Padmavathi (2016) reported bands size ranging from 100 to 400 bp, and Patel et al. (2009) recorded 59 polymorphic bands out of 71 total number bands. The number of polymorphic bands ranged from 2 to 10, with an average of 5.31. A representation of SSR amplification is given in supplementary Fig. 1. However, Yu et al. (2008) reported 3.64 polymorphic alleles and Mishra et al. (2013) noticed 2.31 polymorphic bands. The average polymorphism was 98.46%. Slightly low level of polymorphism noticed by Parekh et al. (2016; 76%) and Dahab et al. (2016; 57.50%) was recorded. But Manonmani et al. (2019) reported as low level of polymorphism as 2.55%. PIC value indicates the informativeness of a marker. PIC values ranged from 0.40 (CES21) to 0.91 (CES28) with an average of 0.61. The range of average PIC was almost similar in the reports of Parekh et al. (2016; 0.11 to 0.74) and Patel et al. (2009; range = 0.91 to 1.00, mean = 0.96). The resolving power of primer was calculated by using the proportion of cultivars containing band. The Rp and PIC give an idea about the primer which could best distinguish the cultivar. Resolving power ranged from 1.58 (COTG28) to 3.96 (CES7), with an average value of 2.37. Mean resolving power ranged from 0.221 (COTG9) to 1.00 (COTG25) with an average of 0.54. The primer index for these 13 primers ranged from 0.55 (CES 21) to 2.82 (CES 28), with a mean primer index value of 1.29. The MI is an overall measure of the effectiveness to identify polymorphism. The usefulness of any marker is established through a balance between the polymorphism level detect by it and its ability to ascertain multiple polymorphisms (Powell et al., 1996). Marker index for SSR primers was observed to be 17.81 (Table 4). Fraction of polymorphism is the relative measure of polymorphic bands to a total number of amplified bands from the set of primers used for the study; in the present study fraction of polymorphism was calculated to be 0.99 suggests higher collective polymorphism. Multiplex ratio reveals the average number of bands generated by each primer. In this study, multiplex ratio was found 5.83 per pair of primers. The effective multiplex ratio is the product of multiplex ratio and fraction polymorphism; in this study, EMR was found 5.31. TB: Total Number of Bands; PB: Number Polymorphic Bands; PI: Primer index; PIC: Polymorphic Information Content; RP: Resolving Power. TP: Total Number of primers; TB: Total Number of Bands; PB: Number Polymorphic Bands; FP: Fractionation of Polymorphism; Hav: Average PIC; MR: Multiplex Ratio; EMR: Effective Multiplex Ratio; MI: Marker Index.
Maker
Band size (bp)
TB
PB
Polymorphism (%)
PI
PIC
RP
Mean RP
SSR
CES7
147–312
5
4
80.00
1.24
0.65
3.96
0.79
CES8
163–432
10
10
100.00
1.92
0.75
3.08
0.31
CES21
94–152
3
3
100.00
0.55
0.40
2.62
0.87
CES28
94–163
10
10
100.00
2.81
0.91
3.29
0.33
COTG9
172–231
10
10
100.00
1.70
0.79
2.21
0.22
COTG25
182–185
2
2
100.00
0.83
0.41
2.00
1.00
COTG28
244–252
6
6
100.00
1.23
0.72
1.58
0.26
HAU216
305–319
4
4
100.00
1.00
0.50
2.00
0.50
HAU218
228–254
6
6
100.00
1.49
0.74
2.00
0.33
HAU225
157–165
4
4
100.00
1.07
0.54
2.00
0.50
MGHES15
191–205
4
4
100.00
0.83
0.41
2.00
0.50
NAU1073
187–194
3
3
100.00
1.25
0.62
2.00
0.67
JESPR247
155–174
3
3
100.00
0.83
0.42
2.08
0.69
Average
5.38
5.31
98.46
1.29
0.61
2.37
0.54
ISSR
HB 08
450–1357
10
9
90.00
2.33
0.81
7.42
0.74
HB 10
313–1292
26
26
100.00
6.56
0.94
9.25
0.36
UBC 807
304–771
9
9
100.00
3.18
0.84
6.08
0.67
UBC 811
312–828
14
14
100.00
5.27
0.91
9.12
0.65
17,898B
85–150
4
3
75.00
1.01
0.70
1.67
0.56
Average
12.60
12.20
93.00
3.67
0.84
6.71
0.60
Marker
TP
TB
PB
FP
Hav
MR
EMR
MI
SSR
13
70
69
0.99
3.36
5.38
5.31
17.81
ISSR
5
63
61
0.97
1.55
12.60
12.20
18.95
3.4.2 ISSR marker
A total of 10 ISSR primers were screened for two genotypes and out of which 5 (50%) primers showed polymorphism. The 5 ISSR primers generated 61 alleles with band size ranging from 85 (17898B) to 1357 bp (HB 08). The number of polymorphic bands ranged from 3 (17898B) to 26 (HB 10), with an average of 12.20. A representation of ISSR amplification is given in supplementary Fig. 2. Kahodariya et al. (2015) reported a 9.44 average number of polymorphic alleles. The percentage polymorphism ranged from 75% (17898-B) to 100% (HB-08, HB-10, UBC-809 and UBC-811) with an average of 95%. Similar observation was recorded by Sethiet al. (2016; 94.30%), Dongre et al. (2007; 88.00%) and Ghuge et al. (2018; 100%). PIC values ranged from 0.70 (17898-B) to 0.94 (HB-10), with an average of 0.839. Kahodariya et al. (2015) reported a similar PIC value. Whereas Sethi et al. (2016) reported slightly lower values. The mean primer index of five ISSR primers was 3.67 with the highest value of PI recovered from HB 10 (6.56) and the lowest value from 17,898 B (1.02). Resolving power ranged from 1.67 (17898B) to 9.25 (HB 10), with an average value of 6.71. Mean resolving power ranged from 0.36 (HB 10) to 0.74 (HB 08) with an average of 0.60. Kahodariya et al. (2015) reported 5.29 mean RP value. Fraction of polymorphism, multiplex ratio, effective multiplex ratio and marker index for ISSR primers were observed to be 0.97, 12.60, 12.20, and 18.95, respectively (Table 4). Kahodariya et al. (2015) reported 7.35 marker index value, 8.62 EMR value. The marker index value recorded by Sethi et al. (2016) was found to be 5.28. In the present study, SSR marker out-performed the ISSR markers exhibiting higher polymorphism (SSR: 98.46%; ISSR 93.00%). Both the markers were found to be highly informative and reproducible in genetic diversity among the selected G. herbaceum genotypes.
3.4.3 Inter-genotype genetic relationship and mantal test
The dendrogram generated from pooled data obtained from SSR and ISSR markers based on Jaccard’s similarity matrix grouped forty-eight desi cotton genotypes into four main clusters: cluster I, cluster II, cluster III and cluster IV with 1, 16, 23 and 8 genotypes, respectively (Fig. 2). Cluster I have only one genotype (4282), cluster II is further divided into two sub-clusters A1 with six genotypes and A2 with eight genotypes. Cluster III is further divided into two sub-clusters namely B1 and B2, having twenty-one. Cluster IV has been divided into two groups C1 and C2, with six and two genotypes, respectively. In the earlier report by Sethi et al. (2016) 65 genotypes were grouped into five main clusters using pooled analysis from SSR and ISSR markers. Likewise, Kahodariya et al. (2015) divided 15 genotypes into seven clusters clearly dividing the old world and new world cotton. A contrast result was observed by Dongre et al. (2007) reported that nineteen genotypes into two groups clearly dividing two species. The genetic coefficient of similarity among the genotypes ranged from 0.15 to 0.70 and the average similarity coefficient was 0.32. The highest genetic distance (0.852) was found between Dhumad and GVhv149, indicating this pair of genotypes highly differed at the genomic level and can be exploited to develop bi-parental mapping population and cotton improvement. The least genetic distance (0.30) was found between 236 and Cross-16, indicating that these genotypes share a common genome. To determine the correlation between marker-based genetic distances with phenotypic distance matrix, Mantel’s correlation test statistics (Z) was applied. The present revealed a low correlation between phenotypic traits and marker-based matrices. The simple correlation between both the genetic matrices was low (r = 0.09). Despite low correlation, both the matrices placed a good number of genotypes in the same clusters in their respective dendrograms. Quantitative traits studied in this investigation were influenced by environmental factors and their phenotype is a product of interaction between genotype and environmental. Furthermore, plants that are morphologically similar need not be genetically similar, since different genetic bases can result in similar phenotypic expression. The low correlation could also be because a large portion of variation detected by molecular markers is non-adaptive and is, therefore, not subject to either natural or artificial selection as compared with phenotypic characters, which in addition to selection pressure are influenced by the environment (Kumar et al., 2013).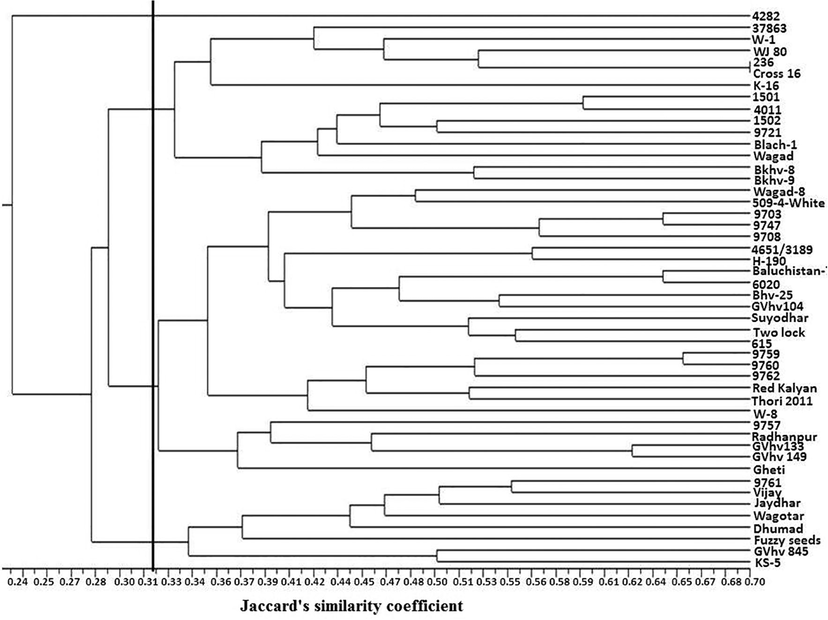
DNA marker based dendrogram showing clustering of 48 cotton genotypes.
4 Conclusion
From current study, it can be concluded that studied genotypes were not only diverse, but also showed heritable variations for studied traits. From the morphological observation it can be concluded that Radhanpur genotype which showed higher seed cotton yield and lint yield; GVhv 845 exhibited good fibre quality parameters can be directly selected as a parent in biparental mating design. Wagad, a popular genotype of western India, exhibited higher oil and comparably less gossypol content. Diversity was also reflected in molecular marker analysis. Both SSR and ISSR techniques, along with proper statistical tools could be successfully applied for genetic diversity study in G. herbaceum. The recorded diversity will be helpful for selecting parents like GVhv-845 for higher fibre strength (26.10 g/tex), Wagad (29.05%) for oil and W8 (51.67%) for seed protein to develop mapping populations and QTL mapping.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Anonymous, 1995.Glanded and glandless cottons. CICR Technical bulletin No.12., ICAR-Central Institute for Cotton Research Nagpurpp 2.
- AOAC (Association of Official Analytical Chemists) Official methods of analysis (10th Edi.), 1965 Washington, DC.
- Natural products (secondary metabolites) Biochem. Mol. Biol. Plants. 2000;24:1250-1319.
- [Google Scholar]
- Linkage disequilibrium and association mapping of drought tolerance in cotton (Gossypium hirsutumL.) germplasm population from diverse regions of Pakistan. Afr. J. Biotech.. 2016;15(46):2603-2612.
- [Google Scholar]
- Genetic divergence in desi cotton (Gossypium arboreum L.). India: Acharya NG Ranga Agricultural University; 2011. Doctoral dissertation
- Variability, heritability and genetic advance in upland cotton (Gossypiumhirsutum L.) Afr. J. Plant Sci.. 2014;8(1):1-5.
- [Google Scholar]
- Genetic diversity in tetraploid and diploid cotton (Gossypium spp.) using ISSR and microsatellite DNA markers. Indian. J. Biotech.. 2007;6(3):349-353.
- [Google Scholar]
- Preparation of enantiomeric gossypol by crystallization. Chirality. 2003;15(6):486-493.
- [Google Scholar]
- Genetic variability, correlation and path analysis among different traits in desi cotton (Gossypium arboreum L.) Afr. J. Agric. Res.. 2014;9(29):2278-2286.
- [Google Scholar]
- Identification of cotton (Gossypium spp.) genotypes by single sequence repeat (SSR) Markers. Int. J. Adv. Multidiscip. Res.. 2016;3(11):62-69.
- [Google Scholar]
- A revised taxonomic interpretation of Gossypium L. (Malvaceae) Rheedea. 1992;2:108-165.
- [Google Scholar]
- Fryxell, P.A. 1979. Genus Gossypium Linnaeus.In: Fryxell P. A. (ed.) The natural history of the cotton tribe. Texas A&M University Press, College Station, Texas, pp. 37-7 1.
- Genome-wide association study of yield components and fibre quality traits in a cotton germplasm diversity panel. Euphytica. 2017;213(3):1-22.
- [Google Scholar]
- Molecular characterisation of cotton genotypes using SSR, ISSR and RAPD markers in relation to fibrequality traits. J. Cotton Res. Dev.. 2018;32(1):1-12.
- [Google Scholar]
- Silver staining of DNA in polyacrylamide gels: linearity and effect of fragment size. Electrophoresis. 1982;3(1):24-26.
- [Google Scholar]
- Assessment of genetic diversity in old world and new world cotton cultivars using RAPD and ISSR markers. Indian J. Biotech.. 2015;14:511-517.
- [Google Scholar]
- Determination of total gossypol and free gossypol content in different varieties of Bt and Non Bt Cotton seed extracts by high-performance liquid chromatography (HPLC) Res. J. Biotech.. 2016;11(2):70-74.
- [Google Scholar]
- Karivaradaraaju, 2007. Processing of cotton seed at Southern India Mills Association Cotton Development and Research Association (SIMA CD&RA).Model training course of cultivation of long staple cotton (ELS).Central Institute for Cotton Research Station, Coimbtore, India. Dec 15-22. pp 74-79.
- The Worldwide Gene Pools of Gossypium arboreum L. and G. herbaceum L., and Their Improvement. In: Paterson A.H., ed. Genetics and Genomics of Cotton. Plant Genetics and Genomics: Crops and Models. New York: Springer; 2009. p. :69-97.
- [Google Scholar]
- Combining molecular-marker and chemical analysis of Capparis decidua (Capparaceae) in the Thar Desert of Western Rajasthan (India) Rev. Biol. Trop.. 2013;61(1):311-320.
- [Google Scholar]
- Molecular diversity studies in cotton (Gossypium hirsutum L.) using SSR markers. Int. J. Curr. Microbiol. App. Sci. 2019;8(7):1731-1737.
- [Google Scholar]
- Validation of fibre quality linked SSR markers derived from allotetraploid (Gossypium hirsutum) in diploid (Gossypium arboreum) Int. J. Sci. Res. Knowl.. 2013;1(9):349.
- [Google Scholar]
- Physical properties of plant and animal materials. New York: Gordon and Breach Science Publishers; 1970. p. :51-83.
- Narayanan, S., Vidyasagar, P., Babu, K. S., 2014, Cotton Germplasm in India — New Trends', In I.Y. Abdurakhmonov (ed.), World Cotton Germplasm Resources, IntechOpen, London. 10.5772/58622,pp 97-118.
- Study on relationship between oil, protein, and gossypol in cottonseed kernels. J. Am. Oil Chem. Soc.. 1975;52(8):312-315.
- [Google Scholar]
- Transcriptomic profiling of developing fiber in levant cotton (Gossypium herbaceum L.) Funct. Integr. Genomics. 2018;18:211-223.
- [Google Scholar]
- Development and validation of novel fibre relevant dbEST–SSR markers and their utility in revealing genetic diversity in diploid cotton (Gossypium herbaceum and G. arboreum) Ind. Crops Prod.. 2016;83:620-629.
- [Google Scholar]
- Genetic diversity analysis in some elite desi cotton cultivars of Gossypium herbaceum and G. arboreum and genetic purity testing of their hybrids through microsatellite markers. Int. J. Plant Sci.. 2009;4(2):464-470.
- [Google Scholar]
- Detection of genetic variation in Ocimum species using RAPD and ISSR markers. 3Biotech. 2015;5:697-707.
- [Google Scholar]
- Assessment of desi cotton (G. herbaceum) genotypes for per se performance under rainfed areas. Adv. Life Sci.. 2016;5(3):865-868.
- [Google Scholar]
- Microscopy techniques and image analysis for evaluation of some chemical and physical properties and morphological features for seeds of the castor oil plant (Ricinus communis) Ind. Crops Prod.. 2011;34(1):1057-1065.
- [Google Scholar]
- Polymorphism revealed by simple sequence repeats. Trends in Plant Sci.. 1996;1(7):215-222.
- [Google Scholar]
- Prakash, A. H (2020) ICAR-AICRP (Cotton) Annual Report (2019-20) ICAR – All India Coordinated Research Project on Cotton, Coimbatore, Tamil Nadu, India - 641 003. Pp A-4.
- Pundir, S. R., 2019. Genetic divergence analysis for morphological and biochemical traits in desi cotton (Gossypium arboreumL.). Doctoral thesis. CCS Haryana Agricultural University, India.
- Variability for gossypol glands in upland cotton (Gossypium hirsutum L.) Source Adv. Plant Sci.. 1991;4:165-170.
- [Google Scholar]
- Properties of cotton seed in relation to design of a pneumatic seed metering device. Indian J. Dryland Agric. Res. Dev. 2015;30(1):69-76.
- [Google Scholar]
- Studies of genetic parameters for seed cotton yield and its contributing characters in Gossypium arboreum L. J. Cotton Res. Dev.. 2014;28:227-229.
- [Google Scholar]
- Rohlf, F.J., 2005. NTSYS-pc: numerical taxonomy and multivariate analysis system version 2.2. Setauket.
- Morphological and molecular diversity patterns in castor germplasm accessions. Ind. Crops Prod.. 2017;97:316-323.
- [Google Scholar]
- Morphological and molecular characterization of desi cotton (Gossypium herbaceum L.) landraces collected from different states of India. Electron. J. Plant Breed.. 2021;12:142-150.
- [Google Scholar]
- Modifying gossypol in cotton (Gossypiumhirsutum L.): A cost effective method for small seed samples. J. Cotton Sci.. 2008;12:202-209.
- [Google Scholar]
- Simple sequence repeats (SSR) and interspersed sequence repeats (ISSR) markers for genetic diversity analysis among selected genotypes of Gossypium arboreum race bengalense. Afr. J. Biotech.. 2016;15(1):7-19.
- [Google Scholar]
- Considering white gold, cotton, for its fiber, seed oil, traditional and modern health benefits. J. Biol. Environ. Sci.. 2020;14(40):25-39.
- [Google Scholar]
- Diversity analysis based on agro-morphological traits and microsatellite based markers in global germplam collections of roselle (Hibiscus sabdariffa L.) Ind. Crop. Prod.. 2017;89:303-315.
- [Google Scholar]
- The impact of seed size and chemical composition on seedling vigour, yield, and fibrequality of cotton in five production environments. Field Crops Res.. 2016;193:186-195.
- [Google Scholar]
- Study on some engineering properties of cotton seeds in relation to the development of a tractor operated seed dibbler. J. Pharmacog. Phytochem.. 2020;9(5):1836-1839.
- [Google Scholar]
- Association analysis of yield and fibre quality traits in Gossypium barbadense with SSRs and SRAPs. Genet Mol Res. 2013;12(3):3353-3362.
- [Google Scholar]
- Genetic evaluation of EST-SSRs derived from Gossypiumherbaceum. Acta Agron. Sin.. 2008;34(12):2085-2091.
- [Google Scholar]
- Associations among lint yield, yield components, and fibre properties in an introgressed population of cotton. Crop Sci.. 2009;49(5):1647-1654.
- [Google Scholar]
- Recent Advances in Cotton Genomics. Int. J. Plant Genom; 2008. Article ID 742304
Appendix A
Supplementary data
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jksus.2022.102336.
Appendix A
Supplementary data
The following are the Supplementary data to this article:







