Translate this page into:
The discovery and simulation analysis of a novel mutation c.40 G < T (V14F) in the NRAS gene in patients with colorectal cancer in Saudi Arabia
⁎Corresponding author. mrahmed1@kau.edu.sa (Mahmood Rasool)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
Background
Colorectal cancer (CRC) is the most diagnosed cancer in men and the second most common cancer in women and remains associated with high morbidity and mortality in Saudi Arabia. The current understanding of genetic heterogeneity of CRC biology encourages the identification of the genetic causes of CRC in the Saudi population.
Methods
In this study, we obtained 89 CRC patients’ tumor samples from Saudi Arabia and investigated the molecular alterations of the NRAS proto-oncogene, GTPase (NRAS) gene in the collected CRC tumor tissue samples to identify gene mutations using DNA sequencing using an automated DNA sequencer ABI 3730xl. The impact of mutations was analyzed using different bioinformatics tools including SwissModel, Missense3D, molecular dynamics simulations using YASARA DYNAMICS and Protein Variation Effect Analyzer (PROVEAN) tool.
Results
We identified a novel mutation c.40 G > T, in one patient in whom valine was replaced by phenylalanine (V14F). Notably, we also identified another mutation in the same codon c.40 G > A where valine is replaced by isoleucine (V14I). Our in-silico analysis revealed that this novel mutation alters the binding affinity of the NRAS gene substantially, and as a result, could have lethal consequences on the downstream signaling genes and pathways including MAPK and PI3K involved in regulating CRC growth and progression.
Conclusions
These findings provide insights into the molecular etiology of CRC in general and particularly in the Saudi population. Thus, these findings in NRAS mutation testing may also guide further treatment modalities, and more personalized therapy may be optimized.
Keywords
NRAS
V14F
Colorectal cancer
Novel mutation
3D homology modeling
1 Introduction
Colorectal cancer(CRC) is the third most common cancer in humans worldwide and ranks second for the most leading lethal malignancy (Siegel et al., 2019, Arnold et al., 2017). Unfortunately, approximately 20 % of CRC are initially diagnosed when they have already metastasized to other organs (Arnold et al., 2013). According to the World Health Organization (WHO) statistics the CRC is the third leading cause of death in Saudi Arabia aged less than 70 years of age group and is responsible for one in eight deaths due to the CRC related malignancies (Bray et al., 2018, WHO 2014).
The study conducted by Peeters et al., on five-year survival of CRC patients demonstrated that the possibility of survival with stage I is 90 % as compared to stage IV patients who showed only 10 % survival, and thus overall five-year survival appears to depend on the CRC stage (Peeters et al., 2015). Notably, the Saudi CRC patient’s overall survival (OS) rate was lower than the internationally reported survival rates based on tumor stage (WHO, 2010).
Despite the availability of various treatment modalities to attenuate CRC development and progression, early diagnosis and correct identification of cancer stages appear to be the most effective strategies to reduce mortality rates, alteration in bowel movements, sudden weight loss, abdominal and pelvic pain, and anemic conditions (Ahnen et al., 2014). These symptoms are considered key indicators for clinicians to start an early and thorough investigation to rule out any possibility of CRC; moreover, patients must also be educated to take such symptoms seriously and report clinicians to be diagnosed perfectly in earlier stages of the disease (Lucente and Polansky, 2018).
From this perspective, there is an indispensable need to implement effective and accurate early-stage screening and monitoring programs to detect malignancies. The adaptation of reliable diagnostic tools of prognostic significance will not only enable the diagnosis of the disease as early as possible but can also be beneficial in predicting chemotherapeutic response outcomes effectively in CRC patients. The in depth understanding and advent of modern molecular tools and techniques have led us to further understand the etiology of cancers at the molecular level. The deep advancement in the knowledge of cancer development and progression has further enhanced our understanding of cancer biology at the molecular level, especially the genetic and epigenetic events that are involved in tumorigenesis; thus, it enables researchers to focus on key signaling pathways involved in the development of cancer, which may be beneficial to reveal novel targets with therapeutic significance (Clarke and Kopetz, 2015).
Notably, the role of RAS family genes in carcinogenesis has been well investigated in CRC, and its oncogenic potential often arises due to the alteration of genes that play a significant role as key mediators as well as in inducing early tumorigenesis in these models. Numerous studies have reported that KRAS proto-oncogene, GTPase (KRAS) and NRAS mutations seem to arise early in the process of colorectal carcinogenesis and thus exert their effect on regulating constitutive signaling and activation of downstream effectors such as mitogen-activated protein kinases (MAPK) and phosphoinositide 3-kinases (PI3K) dependent pathways (Yoon et al., 2014, Ebi et al., 2011), and hence, act as a significant contributor to cancer progression along with other altered genes (Barbie et al., 2009). Multiple studies have demonstrated that, KRAS and NRAS mutations account for 45 % and 5–8 % of CRC cases respectively (Rodriguez-Salas et al., 2017). The majority of mutations in KRAS and NRAS were mainly reported in codons 12 and 13 (90 %) as compared to other codons such as 59, 61, 117, and 146, where they occur less frequently in both KRAS and NRAS genes. However, among the RAS gene family NRAS is the foremost oncogene identified to play a role in melanoma and these NRAS mutations account for almost 20 % of all melanomas (Cancer Genome Atlas Network, 2015). Frequent NRAS mutations were primarily reported at position 61 where amino acid substitution occurs from glutamine (Q) to arginine (R), lysine (K) or leucine (L) (Fedorenko et al., 2013). These mutations have been found to be involved in blocking NRAS into a GTP-bound state, leading to impaired GTPase activity (Simanshu et al., 2017). Furthermore, a deleterious mutation, glycine (G) to aspartic acid (D), has been reported at codons 12 or 13 and comprises 20 % of all NRAS mutations in the amino acid change form (Fedorenko et al., 2013). These functional mutations have been demonstrated to be involved in preventing the association of GAPase-activating proteins (GAP) with NRAS and are more frequently observed in mucosal melanoma than in cutaneous melanoma (Simanshu et al., 2017). Once the NRAS gene is mutated, cascade events occur in the cell signaling pathway, and a series of activations occurs on MAPK signaling as compared to mutations in PI3K–AKT pathways, where they hardly exert such effects on MAPK, indicating that NRAS could be involved in modulating this pathway effectively (Hodis et al., 2012). Alterations in these pathways lead to subsequent dysregulation of the cell cycle and cell proliferation signals (Mendoza et al., 2011).
Hence, in this context, the aim of this research was to study the NRAS gene at the molecular level and define the genetic alterations in the Saudi population to determine the genetic heterogeneity associated with CRC malignancies and better understand the disease manifestation. In this study, we analyzed mutations in the NRAS gene in a Saudi population using automated DNA sequencing and found a novel mutation associated with CRC. At codon 14 valine is substituted with Phenylalanine (V14F) in the NRAS protein. We identified a previously reported V14I mutation in another patient. To understand the functional consequences of the newly identified novel mutation, we carried out in silico modeling studies to establish its impact on cell signaling, especially NRAS downstream effector molecules involved in tumorigenesis.
2 Materials and Methods
2.1 Sample collection and DNA extraction
In this study 89 tumor samples were collected from patients with CRC undergoing surgery at King Abdulaziz University (KAU), Jeddah. Before collecting the tumor and control samples, ethical approval was obtained from the ethical committee of the Center of Excellence in Genomic Medicine Research (CEGMR), KAU. Written Informed consent was obtained from all the recruited patients included in this study. The clinical history of the patients was collected from their medical records. After collection, tumor samples were immediately transported under refrigerated conditions to the CEGMR biobank for further molecular analysis. Genomic DNA was isolated from all tumor tissues following the protocol provided by the DNeasy® Blood & Tissue Kit (QIAGEN). The quality of the extracted DNA was assessed using agarose gel electrophoresis. The quantity and purity were estimated using NanoDrop® ND- 2000 from Thermo Scientific Inc.
2.2 Sequencing and analysis of NRAS gene
Exons 2 and 3 of the NRAS gene were PCR amplified using exon-2 Forward 5ʹ- AAAGTACTGTAGATGTGGCTC-3ʹ and Reverse 5ʹ- GTGAGAGACAGGATCAGG-3ʹ primers (233 bp) and exon-3 Forward 5ʹ- GCATTGCATTCCCTGTGG-3ʹ and Reverse 5ʹ- TAATATCCGCAAATGACTTGC-3ʹ primers (244 bp) respectively (Schulten et al., 2011). The amplified products were subjected to DNA sequencing using an automated DNA sequencer ABI 3730xl. The generated electropherogram was analyzed using BioEdit Sequence Alignment Editor Version 7.2.5 to read sequencing peaks and interpretation of results.
2.3 Homology modeling of NRAS mutants (V14F and V14I) using Swiss-Model server
Template searches for target sequences (NRAS mutants V14F & V14I) using BLAST and HHblits were performed against the SWISS-MODEL template library (Waterhouse et al., 2018) (SMTL, last updated: 2022-04-13, last PDB version included: 2022-04-08). The target sequences of the NRAS mutants (V14F and V14I) were screened against the primary amino acid sequence contained in SMTL using BLAST. An initial HHblits profile was generated for both mutants (Steinegger et al., 2019), followed by iteration of HHblits against Uniclust30 (Mirdita et al., 2016). The obtained profiles were then matched against all SMTL profiles. NRAS V14F and V14I mutant models were built based on the target template alignment using ProMod3 (Studer et al., 2021). The insertions and deletions were remodeled using fragment libraries. The side chains were subsequently rebuilt. Finally, the geometry of the resulting model was normalized using a force field. The global and per-residue quality of the models was assessed using the QMEAN scoring function (Studer et al., 2021, Studer et al., 2020) and the quaternary structure quality estimate (QSQE) score was calculated based on the method developed by Bertoni et al. 2017 (Bertoni et al., 2017).
2.4 Molecular docking of GTP with NRAS (Wild Type), NRAS V14F and NRAS V14I
The SwissDock web server was used to investigate the binding affinity of GTP (Zinc Entry: 60094177) with wild-type NRAS protein, NRAS V14F and NRAS V14I. In SwissDock, the protein–ligand docking algorithm EADock DSS was used, which generates multiple binding modes centered in a virtual box (local docking) or near docking cavities (blind docking) (Grosdidier et al., 2011). Protein-ligand binding energies were assessed using a CHARMM-based scoring function, and the poses with the lowest energy were selected and clustered (Naseer et al., 2016).
2.5 Molecular dynamics simulation studies
Molecular dynamics simulations (MD) for the complex of NRAS (wild type), NRAS V14F, and NRAS V14I bound to GTP were performed in triplicate using YASARA DYNAMICS (YASARA Biosciences GmbH, Vienna, Austria) and followed the in-silico protocol described previously (Krieger et al., 2014). Briefly, the hydrogen bonding network was optimized (Krieger et al., 2012) to increase solute stability, and pKa prediction was performed to normalize the protonation states of the protein residues at the selected pH of 7.4. At physiological concentrations of 0.9 percent sodium chloride (NaCl), ions with excess Na or Cl were added to neutralize the cell. Energy minimization was performed along with a geometric correction of the structure. Conformational stress was eliminated through ‘a short steepest descent minimization”, as also described in the YASARA-based methodology followed by other authors (Shakil et al., 2021). Simulated annealing was employed with a time step of 2 fs. The simulation was run for 46, 39, and 37.75 ns for the complexes of NRAS (wild type), NRAS V14F, and NRAS V14I bound to GTP, respectively. The simulation used the AMBER14 force field (Maier et al., 2015) for the solute, GAFF2 (Wang et al., 2004) and AM1BCC (Jakalian et al., 2002) for the ligand, and TIP3P for water. The cut-off value for the van der Waals forces was 8 Å (default used by AMBER). The 'Particle Mesh Ewald Algorithm' was used without a cut-off value for electrostatic forces. Based on the previously defined algorithms, integration of the equations for the motions was performed with a multiple time step of 2.5 fs for bound interactions and 5.0 fs for unbound interactions at a temperature of 298 K and a pressure of 1 atm (Krieger and Vriend, 2015). Snapshots were captured every 250 ps. A total of 185, 159 and 152 snapshots were retrieved for the complexes of NRAS (wild type), NRAS V14F, and NRAS V14I bound to GTP, respectively. High-resolution images were obtained for the RMSD plots.
2.6 Mutation analysis of NRAS (Wild Type), NRAS V14F and NRAS V14I
The structural effects of replacing phenylalanine and isoleucine with valine at position 14 of the NRAS protein (PDB ID: 6ZIZ) were investigated using Missense 3D software (Ittisoponpisan et al., 2019). Secondary structure change, buried charge exchange, buried proline insertion, buried H-bond breakage, cavity change, buried/exposed change, allowed phi/psi, disulfide bond breakage, buried charge change, collision, buried hydrophilic residue insertion, buried salt bridge rupture, buried charge insertion, buried glycine replacement, and glycine in a bend were calculated to assess the structural effects of these two mutations in the NRAS protein (Rasool et al., 2021). In addition, the Protein Variation Effect Analyzer (PROVEAN) tool (Choi et al., 2012, Choi et al., 2015), which predicts the effects of the substitution or indel of an amino acid in a protein structure on biological function, was used to evaluate the substitution of phenylalanine and isoleucine with valine at amino acid position 14 in the NRAS protein.
3 Results
To investigate and validate the NRAS gene mutation spectrum and its functional correlation with disease susceptibility and treatment outcome in patients with CRC originating from Saudi Arabia, we first amplified exons 2 and 3 of the NRAS gene with a specific pair of primers. After purification of the amplicon cycle, sequencing was performed, and all 89 tumor samples were subjected to DNA sequencing. We identified a novel NRAS mutation (V14F) in a patient with CRC. The identified mutations are depicted in the representative chromatograms in Fig. 1. At codon 14, valine is replaced by phenylalanine (V14F). Notably, we also identified a previously reported mutation in the same codon, where valine is converted into isoleucine (V14I).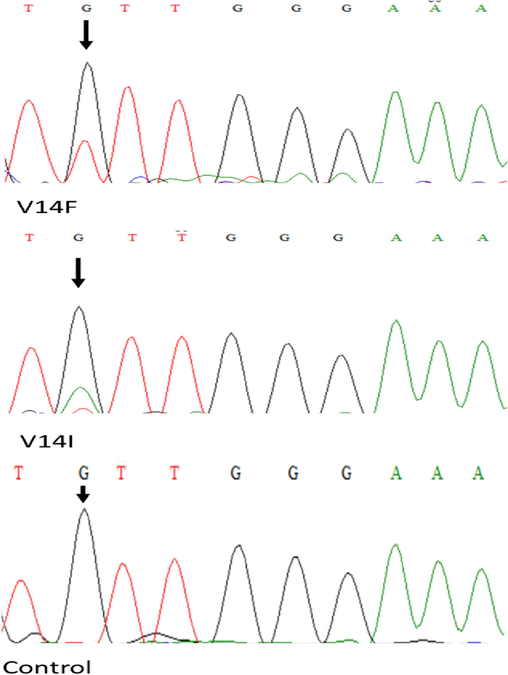
Chromatograms of the two patients with heterozygous mutations c.40 G > T (V14F), c.40G > A (V14I) and control sample.
Homology models for the mutant NRAS proteins (V14F and V14I) were generated using the SWISS-MODEL web server. A total of 11,249 and 11,290 templates were found for NRAS (V14F) and NRAS (V14I), respectively. The crystal structure of the NRAS protein (PDB ID: 6ZIZ) was used as a template to create NRAS (V14F) and NRAS (V14I) homology models (Fig. 2). Docking studies of GTP binding with wild-type NRAS protein (6ZIZ), NRAS V14F and NRAS V14I using the SWISSDOCK web server showed differences in free energy change (kcal/mol) of −14.03, − 5.16 and − 5.80, respectively. The three-dimensional molecular structures of NRAS protein (6ZIZ), NRAS V14F and NRAS V14I bound with GTP, and the two-dimensional structures representing the interaction of amino acids to GTP are shown in Figs. 3, 5 and 7. In the MD simulation experiments (each repeated three times) for the complexes of NRAS (wild type), NRAS V14F, and NRAS V14I with GTP, the contacts (protein–ligand) were plotted against simulation time to understand how the binding forms evolved over the course of the simulation. Hydrogen bonds, hydrophobic contacts and ionic interactions are indicated by red, green and blue dots, respectively. The 'Solute RMSD from the starting structures', namely NRAS WT, NRAS V14F and NRAS V14I with GTP, plotted against the simulation times. The RMSDs for Calpha [RMSDCa], backbone [RMSDBb], and all heavy atoms [RMSDAll] plotted against simulation time for NRAS WT, NRAS V14F, and NRAS V14I with GTP are shown in Figs. 4, 6, and 8, respectively. In addition, analysis of the effects of the mutations on biological function using the PROVEAN web tool revealed that the V14F mutation was deleterious (PROVEAN score: − 4.267), whereas the V14I mutation was classified as neutral (PROVEAN score: − 0.830). However, mutation analysis using Missense 3D software did not reveal any structural changes or damage caused by amino acid substitutions in the NRAS.![NRAS Protein Structure. (A) Wild type NRAS (B) NRAS (V14F), and (C) NRAS (V14I). The amino acid at the 14th position in NRAS (WT), NRAS (V14F), NRAS (V14I) are shown in as space-filling [(Calotte or Corey, Pauling, and Koltun (CPK)] models and visualized using CLC Drug Discovery Workbench 4.0.](/content/185/2024/36/10/img/10.1016_j.jksus.2024.103477-fig2.png)
NRAS Protein Structure. (A) Wild type NRAS (B) NRAS (V14F), and (C) NRAS (V14I). The amino acid at the 14th position in NRAS (WT), NRAS (V14F), NRAS (V14I) are shown in as space-filling [(Calotte or Corey, Pauling, and Koltun (CPK)] models and visualized using CLC Drug Discovery Workbench 4.0.
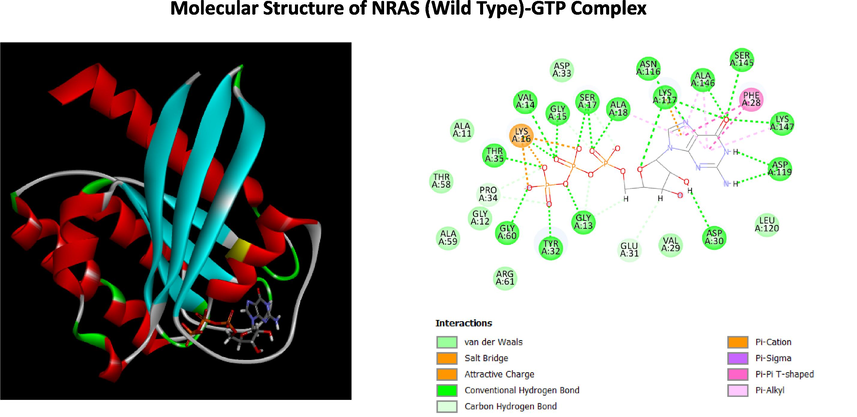
Structure of NRAS-GTP Complex (A) The three-dimensional structure of NRAS (WT) with GTP (B) Two dimensional (2D)-diagram of binding interactions for the complex between GTP and NRAS (WT).
![Solute RMSD from the starting structure (vertical axis) as a function of simulation time. The Calpha [RMSDCa], backbone [RMSDBb], and all-heavy atom [RMSDAll] RMSDs were plotted against the simulation time for NRAS WT interactions with GTP.](/content/185/2024/36/10/img/10.1016_j.jksus.2024.103477-fig4.png)
Solute RMSD from the starting structure (vertical axis) as a function of simulation time. The Calpha [RMSDCa], backbone [RMSDBb], and all-heavy atom [RMSDAll] RMSDs were plotted against the simulation time for NRAS WT interactions with GTP.
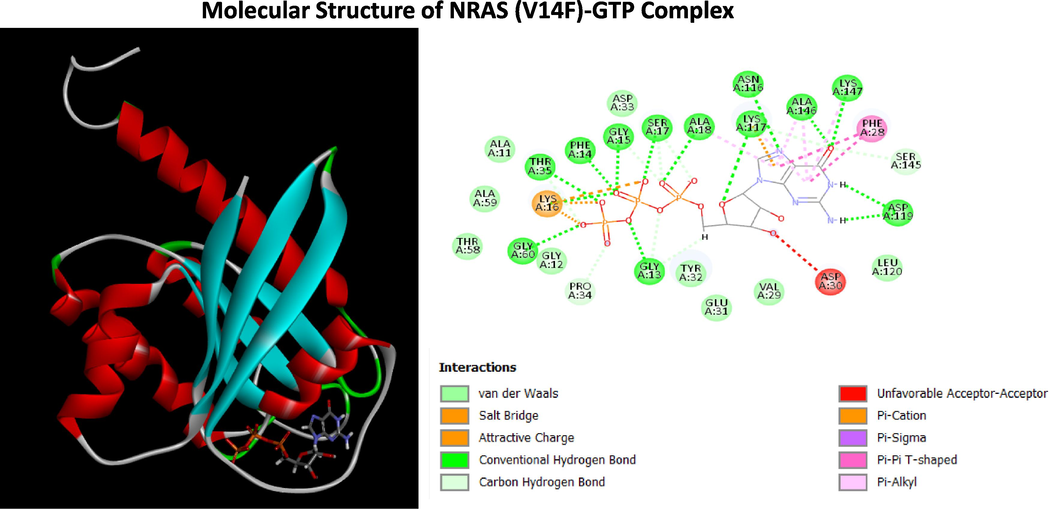
Structure of NRAS (V14F)-GTP Complex (A) The three-dimensional structure of NRAS (V14F) with GTP (B) Two dimensional (2D)-diagram of binding interactions for the complex between GTP and NRAS (V14F).
![Solute RMSD from the starting structure (vertical axis) as a function of simulation time. The Calpha [RMSDCa], backbone [RMSDBb], and all-heavy atom [RMSDAll] RMSDs were plotted against the simulation time for NRAS mutant protein (V14F) interactions with GTP.](/content/185/2024/36/10/img/10.1016_j.jksus.2024.103477-fig6.png)
Solute RMSD from the starting structure (vertical axis) as a function of simulation time. The Calpha [RMSDCa], backbone [RMSDBb], and all-heavy atom [RMSDAll] RMSDs were plotted against the simulation time for NRAS mutant protein (V14F) interactions with GTP.
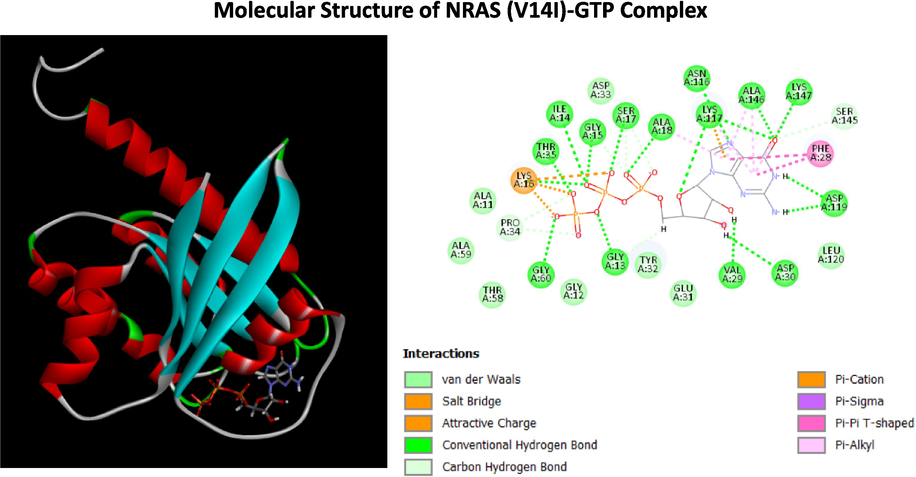
Structure of NRAS (V14I)-GTP Complex (A) The three-dimensional structure of NRAS (V14I) with GTP (B)Two dimensional (2D)-diagram of binding interactions for the complex between GTP and NRAS (V14I).
![Solute RMSD from the starting structure (vertical axis) as a function of simulation time. The Calpha [RMSDCa], backbone [RMSDBb], and all-heavy atom [RMSDAll] RMSDs were plotted against the simulation time for NRAS mutant protein (V14I) interactions with GTP.](/content/185/2024/36/10/img/10.1016_j.jksus.2024.103477-fig8.png)
Solute RMSD from the starting structure (vertical axis) as a function of simulation time. The Calpha [RMSDCa], backbone [RMSDBb], and all-heavy atom [RMSDAll] RMSDs were plotted against the simulation time for NRAS mutant protein (V14I) interactions with GTP.
4 Discussion
Currently, one of the research priorities is to identify the key signalling pathways associated with tumorigenesis, as well as to analyse potential drug resistantmutant phenotype variants in cancer cells. It would be promising for personalized medicine in terms of evaluating and correlating the drug response in individual phenotype variants with distinct sets of genetic alterations might be beneficial in adopting suitable chemotherapeutic strategies effective for the management of such diverse phenotype-related CRC patients (Rasool et al., 2021; Rasool et al., 2024). The recently proposed model by Fearon and Vogelstein for CRC is a multistep genetic process and is considered a benchmark for elucidating the involvement of multiple genes in the development and progression of carcinogenesis, particularly in solid tumours (Fearon and Vogelstein, 1990). According to this model, the initiation of carcinogenesis occurs with the rise of adenomatous polyposis (APC), a tumor suppressor gene that is inactivated and consequently leads to alterations in the KRAS gene. Furthermore, many consequential mutations in the TP53, PIK3CA, and TGF-β pathways that lead to changes in the downstream signalling network are more often correlated with malignant transformation and, as a result, the development and progression of cancer (Samuels and Velculescu, 2004). Although the above-mentioned model described seven distinct mutations that play a vital role in the development and progression of colorectal carcinoma, modern advancements in molecular techniques, especially single-cell sequencing, have revealed more than 80 genes in a single CRC cell (Samuels and Velculescu, 2004). Despite this, recent findings have confirmed the involvement of only 15 genetic mutations as leading drivers associated with carcinogenesis (Wood et al., 2007, Leary et al., 2008).
SwissDock is an integrated in silico tool that can be effectively used for docking simulation studies via a web interface (Grosdidier et al., 2011, Naseer et al., 2016). In this study, we performed docking simulations with the NRAS protein structure (6ZIZ) and mutant structures (V14F and V14I) using the SwissDock web tool. The replacement of valine at amino acid position 14 with phenylalanine or isoleucine in the NRAS protein decreased the binding affinity of these proteins to GTP. Molecular docking studies also confirmed that although both mutations caused structural instability, the V14F interaction with GTP was more unstable than that with V14I. Based on PROEAN and missense 3D web tools, we found that the V14F mutation was deleterious compared to V14I, but neither mutation caused structural damage in the NRAS protein; however, they both functionally damage the protein and resulted in impaired efficiency.
5 Conclusion
Currently, there are no significant efforts to understand the treatment approach for various genetic mutation backgrounds affecting individual CRC patients with altered NRAS genes. Near-term treatment strategies will include combining MEK inhibitors with inhibitors of key intracellular signaling pathways (either involved in RAS signaling or not) or other drugs that disrupt cell cycle checkpoints or epigenetics. To develop personalized therapies, it is important to understand in detail the molecular etiology of each patient. In this study, we investigated the NRAS gene at the molecular level and identified a novel mutation, V14F, in a Saudi patient. This mutation disrupts structural affinity as evaluated using in silico modeling. Further studies with more patient samples and animal models are needed to reach a consensus for therapeutic interventions based on this novel mutatoin in CRC patients.
Disclosure of Funding
This research work was funded by Institutional Fund Projects under grant no (IFPNC-013-141-2020).
CRediT authorship contribution statement
Mahmood Rasool: Writing – original draft, Supervision, Resources, Project administration, Investigation, Data curation. Absarul Haque: Writing – review & editing, Writing – original draft, Conceptualization. Mohammed Alharthi: Investigation, Data curation. Abdulrahman Sibiany: Investigation, Data curation. Mohammed Saad Alamri: Investigation, Data curation. Samer Mohammed Hassan Alqarni: Investigation, Formal analysis. Irfan A. Rather: Validation, Supervision, Project administration. Adeel Gulzar Chaudhary: Visualization, Supervision, Software. Peter Natesan Pushparaj: Writing – review & editing, Resources, Conceptualization. Sajjad Karim: Software, Methodology, Investigation, Formal analysis.
Acknowledgements
This research work was funded by Institutional Fund Projects under grant no (IFPNC-013-141-2020). Therefore, authors gratefully acknowledge technical and financial support from the Ministry of Education and King Abdulaziz University, Jeddah, Saudi Arabia.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- The increasing incidence of young-onset colorectal cancer: a call to action. Mayo Clin. Proc.. 2014;89(2):216-224.
- [CrossRef] [Google Scholar]
- Global patterns and trends in colorectal cancer incidence and mortality. Gut. 2017;66(4):683-691.
- [CrossRef] [Google Scholar]
- Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature. 2009;462(7269):108-112.
- [CrossRef] [Google Scholar]
- Modeling protein quaternary structure of homo- and hetero-oligomers beyond binary interactions by homology. Sci. Rep.. 2017;7(1):10480.
- [CrossRef] [Google Scholar]
- Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin.. 2018;68(6):394-424.
- [CrossRef] [Google Scholar]
- Genomic Classification of Cutaneous Melanoma. Cell. 2015;161(7):1681-1696.
- [CrossRef] [Google Scholar]
- PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics (oxford, England). 2015;31(16):2745-2747.
- [CrossRef] [Google Scholar]
- Predicting the functional effect of amino acid substitutions and indels. PLoS One. 2012;7(10):e46688.
- [Google Scholar]
- BRAF mutant colorectal cancer as a distinct subset of colorectal cancer: clinical characteristics, clinical behavior, and response to targeted therapies. Journal of Gastrointestinal Oncology. 2015;6(6):660-667.
- [CrossRef] [Google Scholar]
- Receptor tyrosine kinases exert dominant control over PI3K signaling in human KRAS mutant colorectal cancers. J. Clin. Invest.. 2011;121(11):4311-4321.
- [CrossRef] [Google Scholar]
- NRAS mutant melanoma: biological behavior and future strategies for therapeutic management. Oncogene. 2013;32(25):3009-3018.
- [CrossRef] [Google Scholar]
- Fast docking using the CHARMM force field with EADock DSS. J. Comput. Chem.. 2011;32(10):2149-2159.
- [CrossRef] [Google Scholar]
- Can Predicted Protein 3D Structures Provide Reliable Insights into whether Missense Variants Are Disease Associated? J. Mol. Biol.. 2019;431(11):2197-2212.
- [CrossRef] [Google Scholar]
- Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J. Comput. Chem.. 2002;23(16):1623-1641.
- [CrossRef] [Google Scholar]
- Assignment of protonation states in proteins and ligands: combining pKa prediction with hydrogen bonding network optimization. Methods in molecular biology (Clifton. N.J.). 2012;819:405-421.
- [CrossRef] [Google Scholar]
- YASARA View - molecular graphics for all devices - from smartphones to workstations. Bioinformatics (oxford, England). 2014;30(20):2981-2982.
- [CrossRef] [Google Scholar]
- New ways to boost molecular dynamics simulations. J. Comput. Chem.. 2015;36(13):996-1007.
- [CrossRef] [Google Scholar]
- Integrated analysis of homozygous deletions, focal amplifications, and sequence alterations in breast and colorectal cancers. Proceedings of the National Academy of Sciences of the United States of America. 2008;105(42):16224-16229.
- [CrossRef] [Google Scholar]
- Colorectal cancer rates are rising in younger adults. JAAPA : Official Journal of the American Academy of Physician Assistants. 2018;31(12):10-11.
- [CrossRef] [Google Scholar]
- ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory Comput.. 2015;11(8):3696-3713.
- [CrossRef] [Google Scholar]
- The Ras-ERK and PI3K-mTOR pathways: cross-talk and compensation. Trends Biochem. Sci. 2011;36(6):320-328.
- [CrossRef] [Google Scholar]
- Uniclust databases of clustered and deeply annotated protein sequences and alignments. Nucleic Acids Res.. 2017;45(D1):D170.
- [CrossRef] [Google Scholar]
- A novel mutation in PGAP2 gene causes developmental delay, intellectual disability, epilepsy and microcephaly in consanguineous Saudi family. J. Neurol. Sci.. 2016;371:121-125.
- [CrossRef] [Google Scholar]
- Peeters, M., Kafatos, G., Taylor, A., Gastanaga, V. M., Oliner, K. S., Hechmati, G., Terwey, J. H., van Krieken, J. H., 2015. Prevalence of RAS mutations and individual variation patterns among patients with metastatic colorectal cancer: A pooled analysis of randomised controlled trials. European journal of cancer (Oxford, England : 1990), 51(13), 1704–1713. doi: 10.1016/j.ejca.2015.05.017.
- Discovery of a novel and a rare Kristen rat sarcoma viral oncogene homolog (KRAS) gene mutation in colorectal cancer patients. Bioengineered. 2021;12(1):5099-5109.
- [CrossRef] [Google Scholar]
- Mutational spectrum of BRAF gene in colorectal cancer patients in Saudi Arabia. Saudi Journal of Biological Sciences. 2021;28(10):5906-5912.
- [CrossRef] [Google Scholar]
- Deciphering gene expression signatures in liver metastasized colorectal cancer in stage IV colorectal cancer patients. Journal of King Saud University - Science. 2024;36(9):103415
- [CrossRef] [Google Scholar]
- Clinical relevance of colorectal cancer molecular subtypes. Crit. Rev. Oncol. Hematol.. 2017;109:9-19.
- [CrossRef] [Google Scholar]
- Oncogenic mutations of PIK3CA in human cancers. Cell Cycle (georgetown, Tex.). 2004;3(10):1221-1224.
- [CrossRef] [Google Scholar]
- Mutational screening of RET, HRAS, KRAS, NRAS, BRAF, AKT1, and CTNNB1 in medullary thyroid carcinoma. Anticancer Res. 2011;31(12):4179-4183.
- [Google Scholar]
- High Throughput Virtual Screening and Molecular Dynamics Simulation for Identifying a Putative Inhibitor of Bacterial CTX-M-15. Antibiotics (basel, Switzerland). 2021;10(5):474.
- [CrossRef] [Google Scholar]
- Siegel, R. L., Miller, K. D., Jemal, A., 2019. Cancer statistics, 2019. CA: a cancer journal for clinicians, 69(1), 7–34. doi: 10.3322/caac.21551.
- RAS Proteins and Their Regulators in Human Disease. Cell. 2017;170(1):17-33.
- [CrossRef] [Google Scholar]
- HH-suite3 for fast remote homology detection and deep protein annotation. BMC Bioinf.. 2019;20(1):473.
- [CrossRef] [Google Scholar]
- QMEANDisCo-distance constraints applied on model quality estimation. Bioinformatics (oxford, England). 2020;36(6):1765-1771.
- [CrossRef] [Google Scholar]
- ProMod3-A versatile homology modelling toolbox. PLoS Comput. Biol.. 2021;17(1):e1008667.
- [Google Scholar]
- Development and testing of a general amber force field. J. Comput. Chem.. 2004;25(9):1157-1174.
- [CrossRef] [Google Scholar]
- SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res.. 2018;46(W1):W296-W303.
- [CrossRef] [Google Scholar]
- Who Strategy for cancer prevention and control in the Eastern Mediterranean Region 2009–2013. 2010.
- WHO. Saudi Arabia Cancer Country Profile. 2014. https://www.emro.who.int/images/stories/ncd/documents/Cancer_Country_Profile_EMR_2014.pdf.
- The genomic landscapes of human breast and colorectal cancers. Science (New York, N.Y.). 2007;318(5853):1108-1113.
- [CrossRef] [Google Scholar]
- KRAS codon 12 and 13 mutations in relation to disease-free survival in BRAF-wild-type stage III colon cancers from an adjuvant chemotherapy trial (N0147 alliance) Clin. Cancer Res. : Off. J. Am. Asso. Cancer Res.. 2014;20(11):3033-3043.
- [CrossRef] [Google Scholar]







