Translate this page into:
In silico identification of novel and conserved microRNAs and targets in peppermint (Mentha piperita) using expressed sequence tags (ESTs)
⁎Corresponding author. s_dhivya@ch.amrita.edu (Dhivya Selvaraj)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Abstract
Objectives
Mentha piperita, generally known as peppermint is a popular herb cultivated in the temperate region around the world belongs to the family Lamiaceae. M. piperita is widely used as traditional food flavoring, in addition they are as well used to treat cold, fever and throat inflammation, and they have anti-viral and anti-fungal properties. The characteristics of MicroRNAs (miRNAs) includes, small size with single stranded non-coding regions having 22 nucleotides. They are evolutionary conserved and actively involved in post-transcriptional regulation of RNAs, which plays various important roles in metabolic and cellular processes of animals and plants.
Methods
The present study reports the potent computational method based on homology search used to find potential miRNA of peppermint. EST sequences were retrieved from NCBI GenBank and BLAST against with plant miRNAs. By various filtration criteria we identified potential miRNAs from miRNA family with suitable properties.
Results and Conclusion
A total of 1216 ESTs were and 1188 unigenes were identified in this study. A set of prospective target genes were identified in peppermint based on sequence homologies mainly encodes for transcription factor, transport inhibitor like protein, GRR1-like protein and DNA-binding protein. The present study endows valuable information on miRNA in peppermint which will accelerate further research in peppermint.
Keywords
miRNA
Computational
BLAST
Target genes
Potential
Peppermint
1 Introduction:
Mint belongs to the family Lamiaceae a largest family of plant with 236 genera and 7000 species (Marzouk et al., 2018) and genus Mentha, consist of 42 species, 15 hybrids and subspecies, cultivars and varieties (Kalemba and Synowiec, 2020). Mentha piperita is generally called as peppermint is a perennial herb widely used by greater part of people globally in various forms like leaf, leaf extract oil and leaf water (Ekor, 2014). Due to its rich flavor and fragrance peppermint has economic value both in the domestic and international market and an important export commodity that fetches good foreign revenue. Peppermint is a valuable plant for its distinctive aroma and medicinal value. It is a resource for various chemical substances like terpenoids and phenylpropanoids, having varied chemical structures that are part of particular metabolism (Sangwan et al., 2001). It is also rich in enormous bioactive compounds and contains sugars, saponins, alkaloids, anthraquinones, and quinines (Padmini et al., 2008).The peppermint oil logs for various antispasmodic activities like antivomiting, carminativum, stomachic and antimicrobial properties. The main compounds such as menthol, menthone, and menthofuran which gives unique aroma (da Silva Ramos et al., 2017).
miRNAs are tiny non-coding RNAs, the length is about 22–24 nucleotides in size. A DNA sequence first transcribes to primary miRNAs followed by precursor miRNAs and in turn into mature miRNAs. Mostly, miRNAs associate with the target mRNAs by 3′ UTR to curb its expression (Denli et al., 2004). It also interacts with gene promoters and 5′ UTR coding sequence (Broughton et al., 2016); in addition under specific condition it activates gene expression (Vasudevan, 2012). Besides it shuttles among diverse sub-cellular organelles to control the frequency of transcription and translation (Makarova et al., 2016), involves in differentiation and development process, abiotic and biotic stress response (Ren et al., 2012). With robust computational and experimental approaches several miRNAs were identified in plants (Bartel and Bartel, 2003), and through comparative genomics approach these miRNAs were found to be highly conserved among species. This miRNA can also be identified by the use of expressed sequence tag (EST) and a Genome Survey Sequence (GSS) (Zhang et al., 2007). EST analysis has gained significance when compared with other approaches like conserved miRNAs and be capable of identifying without knowing the whole genome sequences. It offers straight confirmation for miRNA expression than from genomic sequence surveys without any much specified software. By this method several miRNAs are effectively recognized in diverse plant species.
The genome-wide miRNA profiling have been developed using three major platforms like small RNA cloning, microarray and qRT-PCR further by miRNA-Seq which provides single-nucleotide resolution of miRNA species. The deep sequencing approach is yet to be successfully developed. It is a challenge to identify unique miRNAs from the big group of small RNA sequences. Hereby in silico computational approach is employed in identification of novel and conserved miRNAs in peppermint which has immense medicinal and economic importance. In the current research we had recognized novel and conserved miRNAs using expressed sequence tags (ESTs) of peppermint. miRBase is a public miRNA database that has been first deposited by Kozomara et al., (Kozomara and Griffiths-Jones, 2014). In current research, we applied expressed sequence tag-based homology search to discover miRNAs in peppermint by presently available 1216 expressed sequence tags ESTs from NCBI EST database. From the outcomes of our results, only one possible miRNA, from the peppermint family has been identified. This recently recognized miRNA plays an essential function in diverse biological processes. This study reveals the key features of recognized peppermint miRNA.
2 Methods
2.1 Computational reorganization of mint miRNA
In total 74 mature miRNA taken from miRNA sequence database miRBASE Version 21 (https://www.mirbase.org). To find conserved and novel miRNAs in peppermint, 43 distinctive miRNAs were predicted and chosen for next studies. On the whole 1216 EST sequences of peppermint (Mentha piperita) are taken in NCBI dbEST database (https://www.ncbi.nlm.nih.gov/nucest/). Blast 2.2.30 + an offline version of BLAST was used for further studies (ftp://ftp.ncbi.nlm.nih.gov/blast). Nearly 1188 contigs were identified using CD HIT server from which 1061 Unigenes were annotated.
2.2 Expressed sequence tag assembly and functional annotation
The ESTs were assembled employing CLC Genomics Workbench 8.0 (https://www.clcbio.com/products/ clc-genomics-workbench/). Gene ontologies and functional annotation were analyzed by BLAST2GO suite21. BLASTn and BLASTx servers were employed in similarity search against the viridiplantae family. The unigenes were annotated based on similarity search using of e-10.
2.3 Mature miRNA identification
Pair-wise alignment of assembled EST sequences against a unique miRNA database with a threshold value of e-10 and window size of 7 was carried out using the BLAST algorithm using mature miRNAs extracted from the miRBase database. miRNA candidates were selected using homology search approach of the threshold of 18nt length with no gaps. Further study was followed by the known miRNAs were selected that are closely matched with the assembled sequence and subject to BLASTx search against non-redundant protein database. Fig. 1 represents the miRNA scheme and EST analysis in peppermint.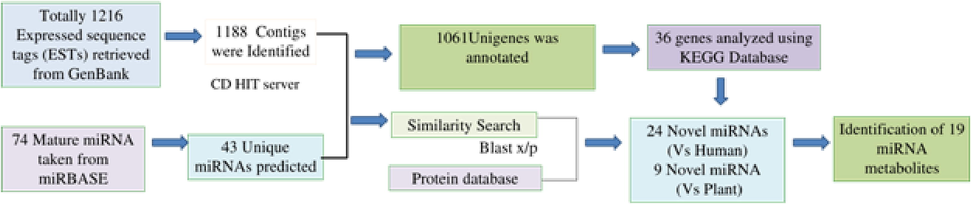
Illustrative representation of the method followed for the identification of miRNAs from mint ESTs.
2.4 miRNA secondary structure prediction
The non-protein coding predecessor sequence was employed to envisage the secondary structure using Mfold software which uses energy minimization method, protein coding sequences were removed prior to prediction (Kozomara and Griffiths-Jones, 2014; Zuker, 2003), filter criteria was followed to select pre-miRNA with the length of 60 nt properly folded to form stem-loop-hairpin secondary structure with maximum of 6 nt mismatches. There must be no breaks allowed between the miRNA/miRNA* duplex, A + U content must be between 30 and 70 percent, and the projected secondary structure must have a higher minimal folding free energy index (MFEI) and negative minimal folding free energy (MFE), which distinguish miRNAs from short RNAs. The negative free was calculated using the fact that:
2.5 miRNA target genes prediction
The important criteria for miRNA target prediction are seed match, free energy, conservation, target site abundance and site accessibility. The target miRNAs were predicted using homology algorithm (Rhoades et al., 2002), psRNA target server was used to find miRNA targets with default parameters. The prospective miRNAs as query are searched against with mRNAs and assembled ESTs of peppermint. Target accessibility was granted maximum energy to unpair the target site (UPE) by adhering to the specifications of maximum exception being 3.0, length of complementarily scoring (hspsize) being 20, and flanking length approximately about the size of target site for accessibility analysis which is 13 bp downstream and 17 bp upstream. The highest mismatch at the complementary site is 4 without gaps, while the choice of the central mismatch resulting to translational inhibition is 9–11 nucleotides.
2.6 Phylogenetic analysis of mature miRNAs
The standalone blast search tool was used against miRBase database and finally matured miRNAs were predicted. Multiple sequence alignment tool ClustalW, used to align sequence of precursor miRNA and newly identified miRNAs. MEGA version 6.25 was used to conduct phylogenetic and molecular evolutionary analyses and phylogenetic tree was built using maximum likelihood method (Olsen et al., 1994).
2.7 Functional annotation and pathway analysis of miRNA target genes
The target miRNA gene sequences were matched to the non-redundant protein sequence database of NCBI GO keywords using the BLAST2GO 3.0 tool (https://balst2go.com/blast2go-pro/download-b2g). The Gene Ontology (GO) database was used to look into biological processes, cellular components, and molecular functions. The Kyoto Encyclopedia of Genes and Genomes (KEGG) (Kanehisa and Goto, 2000) KASS server was used to annotate target miRNAs, its metabolic pathways, and their networks (Moriya et al., 2007).
2.8 Structure prediction of key metabolites and its binding pockets of M. Piperata
The 3D structure and active sites of identified key metabolites like (s) -limone 3- monooxygenase, (+) menthofuran synthase, (+) pulegone reductase, (+) neomenthol dehyrogenase and vestitone reductase were modeled and predicted by RaptorX (https://raptorx.uchicago.edu/BindingSite/) (Källberg et al., 2012).
3 Results
The dbEST database yielded a total of 1216 EST sequences, which were subsequently assembled and functionally annotated. 1188 unigenes were produced by the EST assembly after redundancy was eliminated. For peppermint unigenes, the typical read length was 519 bp. Table 1 displays the summary data for EST assembly, clustering, and annotation.
Item
Number
Total number ESTs
1216
Number of contigs
1188
Total number of bases in unigenes
617,396
Maximum length of contigs
1147
Minimum length of contigs
157
Average length of contigs
519
Number of unigenes annotated
1061
Total number of unannotated unigenes
93
The BLAST2GO suite was used, and unigenes were mapped and assigned to Gene Ontology (GO) keywords of the unigenes shown in Fig. 2 for the similarity search BLASTx against numerous public databases. 1061 (87.25 %) of the 2710 unigenes received significant hits in the similarity search, while 93 (7.6 %) had no significant similarity. One species only, M. piperita, is thought to contain unannotated unigenes. Additionally, the M. piperita unigenes demonstrated the utmost likeness with Musa acuminata, followed by Phoenix dactylifera and Elaeis guineensis.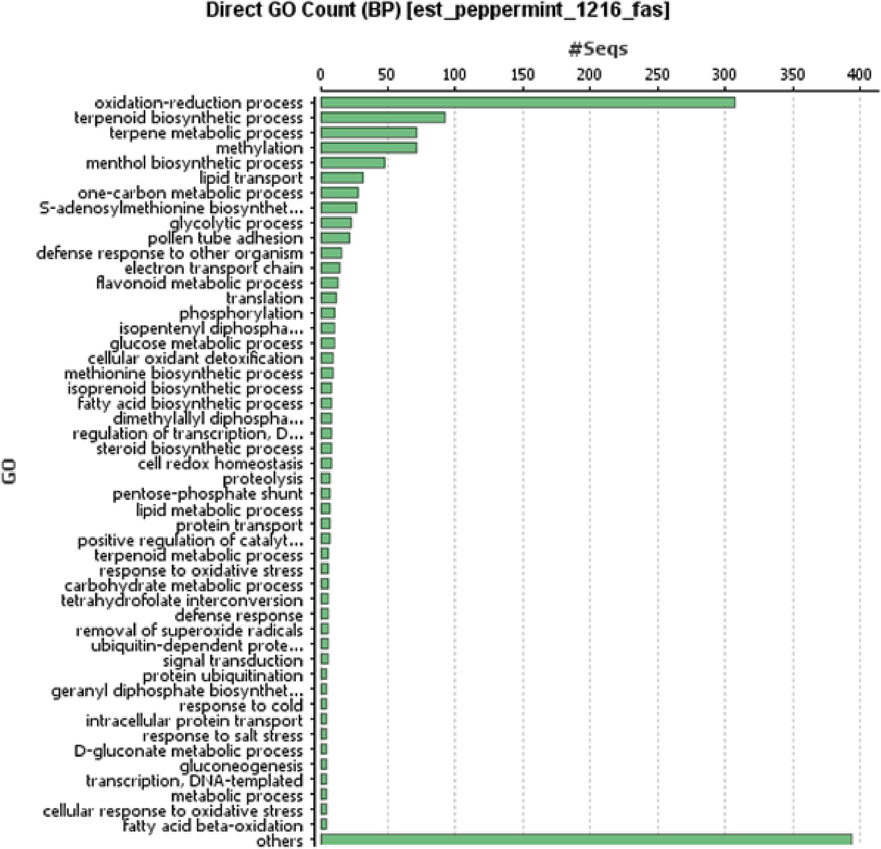
Species distribution of mint unigenes based on sequences similarity search.
Table 2 shows the Gene Ontology (GO) terms that were assigned to unigenes. Further characterization includes oxidation reduction, terpenoid biosynthesis, terpene metabolism, methylation, menthol biosynthesis, lipid transport, one-carbon metabolism, and S-aenosylmethionine production, among other molecular processes. Small molecular binding, transferase activity, and hydrolase activity genes all played an important role in molecular functioning. Genes involved in metabolic, single-organism, cellular, and biological processes have been found. Furthermore, the cell's cellular components had the greatest representation in both cell and organelle.
Source
Homologous miRNA
% of AU of Pre-miRNA
Free energy Kcal/ mol
Predicted miRNA sequence
LM
MN
e- value
miRNA-1
pmtmiR159
50
−50.90
UAUGGCGAGUAGGAAAAUCU
20
0
1.9
miRNA-2
pmtmiR5021
58
−53.33
CAUAUGGACAUCUAGAUGG
19
1
1.7
miRNA-3
pmtmiR772
58
−52.78
UAUGGCGAGUAGGAAAAUC
19
2
1.5
miRNA-4
pmtmiR414
58
−67.50
UCUCUUCUCAUCUC UCAUC
19
3
3.7
miRNA-5
pmtmiR5658
64
−63.70
UGUGAUGAUGAUGAAAC
17
2
3.5
miRNA-6
pmtmiR159
52
−65.25
GAAGAGCUCCUU AGUUCAAGG
21
2
3.5
miRNA-7
pmtmiR8166
58
−44.69
AGUGUAGAAAGUUCUC
17
2
0.41
miRNA-8
pmtmiR156j
52
−27.22
UGACAAAG AGAGAGCAC
17
2
0.41
miRNA-9
pmtmiR5021
77
−40.57
UGAGAAGAAGAA AAAA
18
4
3.2
Peppermint unigene sequences have been matched with Kyoto encyclopaedia of genes and genomes reference pathways (KEGG). On the whole, 36 number of KEGG pathways was recognized with biosynthesis of monoterpene as the mainly ample in the classes of enzyme (+)-menthofuran synthase, (+)-pulegone reductase, (4S)-limonene synthase and vesititone reductase, followed by ATP-dependent degradation of ubiquitinated proteins pathways. This study supported the customary idea about peppermint having medicinal values.
3.1 Identification of miRNA in peppermint ESTs
A total of 1188 unigenes were used to identify miRNAs, with the majority of mature plant miRNAs being evolutionarily conserved. MiRBase (https://www.mirbase.org/cgi-bin/browse.pl) was used to obtain 74 mature miRNAs. After eliminating the duplicated sequences, 43 unique miRNAs were used as a reference, and 1188 unigenes were BLASTed against them using settings such as e-10, word, and window size of 7. With 9 unigenes, the search yielded 24 possible mature miRNA showing homologs. The anticipated miRNA length is between 16 and 22 nucleotides, which is consistent with earlier plant research. Mfold software was used to create secondary structures for selected miRNAs, and various filters were employed to pick candidate miRNAs. Nine miRNAs had higher free energy and content as prospective miRNAs, out of a total of 24. The stability of the hairpin stem loop secondary structure of the precursor miRNA was determined using the terms minimum free energy (MFE) and anticipated value (E-value). Therefore, a lower value of MFE implies that the secondary hairpin stem loop structure of the precursor miRNA is more stable. Fig. 3 provides the anticipated mRNA structures.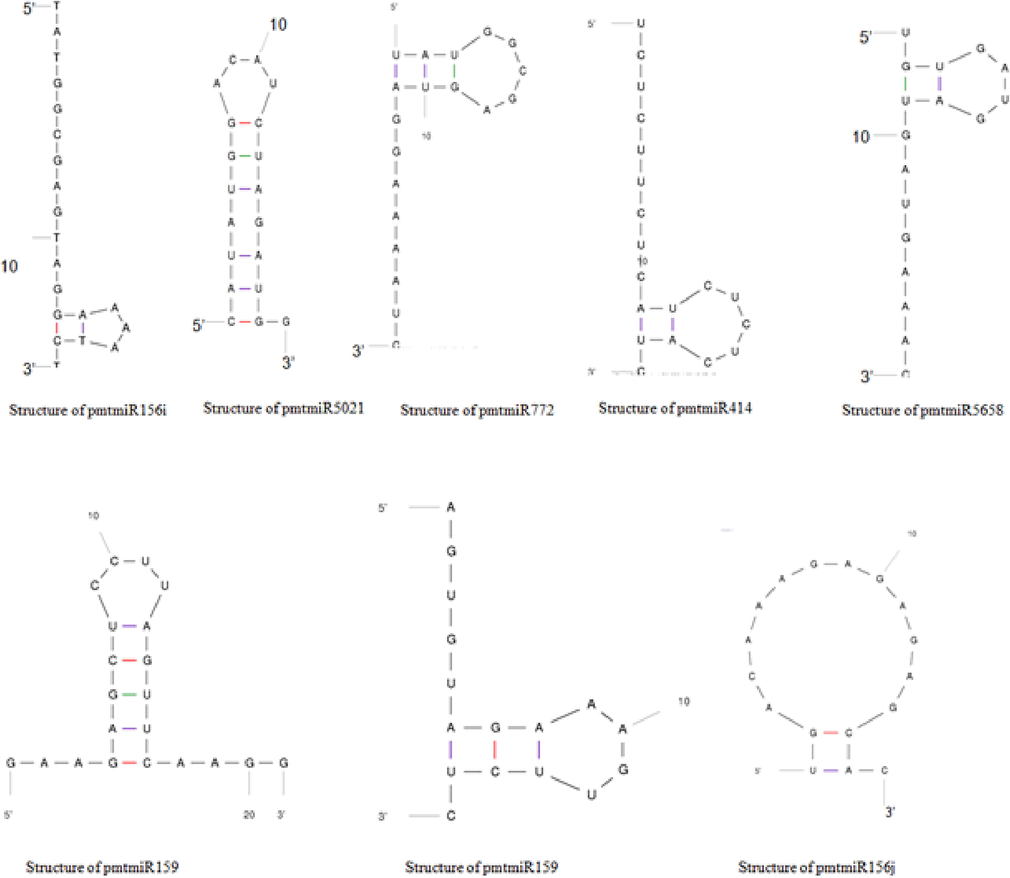
Structures of predicted mRNA.
This study shows the occurrence of nine miRNA from 1216 ESTs of peppermint, according to Zhang, et al 2007 the likelihood of discovering miRNA is one per 1000 ESTs. The nucleotide percentage for recently recognized miRNA in peppermint ESTs are tabulated in Tables 3 and Table 4.
miRNA
Mature miRNA sequence
miRNA family
A%
C%
G%
U%
A/U
G/C
A + U%
miRNA-1
UAUGGCGAGUAGGAAAAUCU
pmtmiR5021
35
10
30
25
1.4
2.5
60
miRNA-2
CAUAUGGACAUCUAGAUGG
pmtmiR5021
31.5
15.8
26.3
26.3
1.2
1.6
58.1
miRNA-3
UAUGGCGAGUAGGAAAAUC
pmtmiR772
36.8
10.52
31.5
21
1.75
3
57.8
miRNA-4
UCUCUUCUCAUCUCUCAUC
pmtmiR414
10.5
42.1
0
47.3
0.2
0
57.8
miRNA-5
UGUGAUGAUGAUGAAAC
pmtmiR5658
35.3
5.88
29.4
29.4
1.2
5
64.7
miRNA-6
GAAGAGCUCCUUAGUUCAAGG
pmtmiR159
28.6
19.0
28.6
23.8
1.2
1.5
52.4
miRNA-7
AGUGUAGAAAGUUCUC
pmtmiR8166
31.3
1.25
2.5
31.3
1
0.5
62.6
miRNA-8
UGACAAAG AGAGAGCAC
pmtmiR156j
47
18.7
29.4
5.88
8
1.7
52.8
miRNA-9
UGAGAAGAAGAAAAAA
pmtmiR5021
68.7
0
25
16
11
4
84.7
S.No
Accession No
Target Protein
miRNAs
Inhibition KO number
GO Annotation Biological process) Functions
1
AW254845.1
histone H1-like
ML1267
K11275
Stabilizes chromatin structure
2
AW254953.1
(S)-limonene 3- monooxygenase
ML1122
K15089
Monoterpene biosynthesis
3
AW255209.1
26S protease regulatory subunit 6A
ML201
T03446
ATP-dependent degradation of ubiquitinated proteins
4
AW255784.1
26S protease regulatory subunit 6A
ML852
K03065
ATP-dependent degradation of ubiquitinated proteins
5
AW255882.1
(+)-menthofuran synthase
ML968
K15093
Monoterpene biosynthesis
6
AW255952.1
(+)-pulegone reductase
MW298
T10000
Monoterpene biosynthesis
7
AW255510.1
glycine-rich protein A3-like
ML536
T05027
Interaction in cell wall associated
Proteins
8
AW254888.1
(4S)-limonene synthase
ML1312
K15088
Biosynthesis of sec metabolite
9
AW255223.1
4-hydroxy-3-methylbut-2-en-
1-yl diphosphate reductase
ML218
K03527
Isoprenoid biosynthesis
10
AW255083.1
(-)-camphene/tricyclene synthase, chloroplastic-like
ML1378
K07385
1,8-cineole synthase
11
AW254725.1
(-)-alpha-terpineol synthase- like
ML1011
T01084
Monoterpenoid biosynthesis
12
AW255955.1
(+)-neomenthol dehydrogenase
ML536
K15095
Monoterpenoid biosynthesis
13
AW255293.1
S-denosylmethionine synthase
ML300
K00789
Methylation in response to stress
14
AW255832.1
trans-resveratrol di-O- methyltransferase
ML900
K16040
Stilbenoid, diarylheptanoid and gingerol biosynthesis
15
AW25593.1
(+)-pulegone reductase
MW188ED
K15092
Monoterpene Biosynthesis
16
AW255630.1
acyl-coenzyme A thioesterase
ML69
K17362
Activates thiolester hydrolase
17
AW255783.1
vestitone reductase
ML851
K13265
Isoflavonoid biosynthesis
18
AW255061.1
membrane- associated progesterone receptor
ML1356
K17278
Progesterone binding
19
AW255186.1
MADS-box transcription factor
ML174
K09264
Cytoskeletal dynamics
3.2 miRNA target prediction
Peppermint contigs/unigenes were analysed for the miRNAs prediction which regulates gene expression and actively involved in growth development, stress stimulus, defense mechanism. psRNA target server, a homology based tool was employed in prediction of peppermint miRNA from target mRNAs. A total of 19 targets for the new identified mature miRNAs of peppermint were reported. Among 19, nine were of miRNAs which are represented in the Table 3.
3.3 Phylogenetic analysis
The anticipated precursor miRNA sequences were aligned using Clustal W, and phylogenetic analysis was utilized to establish the evolutionary link using MEGA's maximum likelihood technique. MW298 and ML1312 were discovered to be closely related among 19 predicted miRNA sequences that are active in Monoterpene biosynthesis, indicating (+)-pulegone reductase and (4S)-limonene synthase, respectively, as illustrated in Fig. 4.Fig. 5..
Phylogenetic tree prepared using maximum likelihood method showing relationship between newly identified pre-miRNAs of mint with their closely related families.
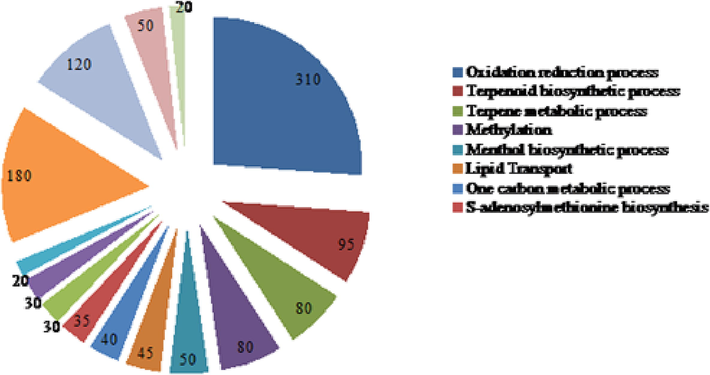
Functional annotations of miRNAs in M.piperata.
3.3.1 KEGG pathway analysis and GO term annotation
To better comprehend the roles of the newly discovered target genes, Gene Ontology (GO) concepts were applied. In terms of biological process, molecular function and cellular components, the targeted genes were assigned to regulatory networks that are active in oxidation reduction process, terpenoid biosynthesis, terpene metabolism, methylation, menthol biosynthesis, lipid transport, one carbon metabolism, S-adenosylmethionine biosynthesis, glycolytic process, pollen tube adhesion defence response, integrated component membrane, and cytoplasm endoplasmic reticulum.
3.4 Structure prediction of key metabolites and its binding pockets of M. Piperata
The binding pocket residues were found in the enzymes (s) -limone 3- monooxygenase (EC:1.14.14.143), (+) menthofuran synthase (EC:1.3.1.81), (+) pulegone reductase (EC:1.3.1.81), (+) neomenthol dehyrogenase (EC:1.3.1.81), and vestitone reductase (EC:1.3.1.81) (EC: 1.1.1.348). a. Binding pocket locations were utilised to assess the quality of an enzyme's predicted structure. It denotes the number of times the selected binding pocket was discovered in a set of important metabolites. From 19 to 23 residues, the number of binding pockets and their multiplicity were changed. Figs. 6 and 7 shows the binding residues of important metabolite structures as well as their positions. The hydrophobicity of the binding residues is roughly 50%.![Modelled structure of key metabolites and binding pockets of M. piperata. (A) Binding sites of (s) -limone 3- monooxygenase [G12 N14 K15 G16 I17 R37 L65 D66 V67 N93 A94 G95 V96 V180 S181 K242 G267 A269 T271 D272 273 T274]. (B) Binding sites of (+) menthofuran synthase [T68 A71 G72 T75 S76 T79 M130 P135 I136 I139 P140 R141 P201 F202 G203 R207 I208 C209 P210 G211 F214 G215 V219]. (C) binding sites of (+) pulegone reductase [G15 T17 G18 F19 L20 I39 R40 L69 M88 A89 H90 P91 L92 T130 S131 S132 Y168 K172 P195 T196 W197 I198 S210]. (D) binding sites of (+) neomenthol dehyrogenase [R99 V115 A116 R128 M292 A295 G296 T299 T300 A303 P359 F360 L363 L364 R366 P426 F427 G428 R432 G433 C434 P435 G436 F439 A440 Y444]. (E) Binding sites of vestitone reductase [P52 M135 T139 G163 S164 V165 A184 G185 K189 Y205 N229 V230 C251 G252 V254 S255 Y257 F281 V282 V283 V326 G329].](/content/185/2023/35/4/img/10.1016_j.jksus.2023.102604-fig6.png)
Modelled structure of key metabolites and binding pockets of M. piperata. (A) Binding sites of (s) -limone 3- monooxygenase [G12 N14 K15 G16 I17 R37 L65 D66 V67 N93 A94 G95 V96 V180 S181 K242 G267 A269 T271 D272 273 T274]. (B) Binding sites of (+) menthofuran synthase [T68 A71 G72 T75 S76 T79 M130 P135 I136 I139 P140 R141 P201 F202 G203 R207 I208 C209 P210 G211 F214 G215 V219]. (C) binding sites of (+) pulegone reductase [G15 T17 G18 F19 L20 I39 R40 L69 M88 A89 H90 P91 L92 T130 S131 S132 Y168 K172 P195 T196 W197 I198 S210]. (D) binding sites of (+) neomenthol dehyrogenase [R99 V115 A116 R128 M292 A295 G296 T299 T300 A303 P359 F360 L363 L364 R366 P426 F427 G428 R432 G433 C434 P435 G436 F439 A440 Y444]. (E) Binding sites of vestitone reductase [P52 M135 T139 G163 S164 V165 A184 G185 K189 Y205 N229 V230 C251 G252 V254 S255 Y257 F281 V282 V283 V326 G329].
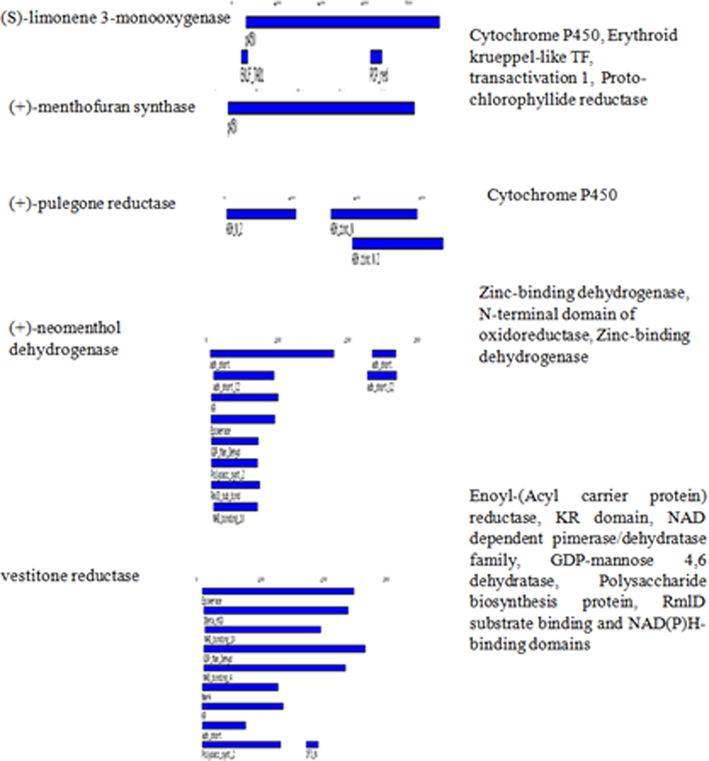
Target motif metabolites gene in M. piperata.
4 Conclusion
The peppermint (Mentha piperita) is known for its medicinal properties. EST sequence of M. piperita was used for miRNA prediction and target identification by computational approach. The present study identified nine novel miRNA and their targeted genes in peppermint that has major significance in metabolic process. The identified 41 target genes modulate a variety of regulatory molecules, including transcription factors, enzymes, and secondary messengers, to control the expression of several genes. By computational approach this study systematically identified the miRNAs and characterizes the genes and their potential regulatory miRNAs for the synthesis of bioactive components such as limonene, menthofuran, pulegone, neomenthol and vestitone of terpene family. Conclusively these finding may further enhance our understanding towards peppermint miRNAs and their importance in biological process.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Acknowledgement
This project was supported by Researchers Supporting Project number (RSPD2023R712), King Saud University, Riyadh, Saudi Arabia.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- MicroRNAs: at the root of plant development? Plant Physiol.. 2003;132(2):709-717.
- [CrossRef] [Google Scholar]
- Pairing beyond the Seed Supports MicroRNA Targeting Specificity. Mol. Cell. 2016;64(2016):320-333.
- [CrossRef] [Google Scholar]
- Chemical Composition and In Vitro Antioxidant, Cytotoxic, Antimicrobial, and Larvicidal Activities of the Essential Oil of Mentha piperita L. (Lamiaceae) Scientific World J. 2017:4927214.
- [CrossRef] [Google Scholar]
- Processing of primary microRNAs by the Microprocessor complex. Nature. 2004;432(7014):231-235.
- [CrossRef] [Google Scholar]
- The growing use of herbal medicines: issues relating to adverse reactions and challenges in monitoring safety. Front. Pharmacol.. 2014;177
- [CrossRef] [Google Scholar]
- Agrobiological Interactions of Essential Oils of Two Menthol Mints: Mentha piperita and Mentha arvensis. Molecules. 2020;25:59.
- [CrossRef] [Google Scholar]
- Template based protein structure modeling using the RaptorX web server. Nat. Protoc.. 2012;7(8):1511-1522.
- [CrossRef] [Google Scholar]
- KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res.. 2000;128(1):27-30.
- [CrossRef] [Google Scholar]
- miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res.. 2014;42(Database issue):D68-D73.
- [CrossRef] [Google Scholar]
- Intracellular and extracellular microRNA: an update on localization and biological role. Prog. Histochem. Cytochem.. 2016;51(3–4):33-49.
- [CrossRef] [Google Scholar]
- Comparative study of Mentha species growing wild in Egypt: LC-ESI-MS analysis and chemosystematic significance. J. Appl. Pharm. Sci.. 2018;8(08):116-122.
- [CrossRef] [Google Scholar]
- KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acid. Res.. 2007;182–185
- [CrossRef] [Google Scholar]
- fastDNAmL: A tool for construction of phylogenetic trees of DNA sequences using maximum likelihood. Computer Applications in the Biosciences. 1994;1994:41-48.
- [CrossRef] [Google Scholar]
- Comparative study on composition and antioxidant properties of mint and black tea extract. Int. J. Food Sci. Technol.. 2008;43:1887-1895.
- [Google Scholar]
- Identification of novel and conserved Populus tomentosa microRNA as components of a response to water stress. Funct. Integr. Genomics.. 2012;12(2):327-339.
- [CrossRef] [Google Scholar]
- Regulation of Essential Oil Production in Plants. In: Plant Growth Regulation. Kluwer Academic Publishers; 2001.
- [Google Scholar]
- Posttranscriptional upregulation by microRNAs. Wiley Interdiscip. Rev. RNA. 2012;3:311-330.
- [CrossRef] [Google Scholar]
- Identification of cotton microRNAs and their targets. Gene. 2007;397(1–2):26-37.
- [CrossRef] [Google Scholar]
- Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res.. 2003;31:3406-3415.
- [CrossRef] [Google Scholar]







