Translate this page into:
Degradation of oil and hydrocarbons derivatives by isolated yeast strains from Ras Tanura in Saudi Arabia
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
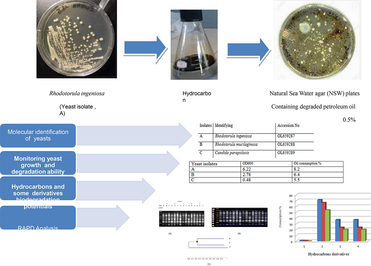
Abstract
Saudi Arabia heavily depends on oil production, as 33.42 % of GDP is generated from the oil sector (Saudi Arabian Monetary Agency, 2019). Due to previous fact, microorganism considers the main concern for oil biodegradation process. As a result of highly advantage of employed yeast for oil biodegradation than bacteria, this study was carried out to isolate crude oil biodegrading yeast from sediments from three points of Ras Tanura City (26°38′38″N and 50°9′33″E), eastern province, Kingdom of Saudi Arabia. Among nine yeast isolates, only three isolates selected due to remarked biodegrading activities. Based on 18S rRNA Partial gene sequence, A, B and C yeast isolates were identified and submitted as Rhodotorula ingeniosa (OL659287), Rhodotorula mucilaginosa (OL659288) and Candida parapsilosis (OL659289) with 100 % identity respectively. A isolate was superior for oil consumption (8.2 %) comparing with B and C isolates which showed 6.4 % and 5.5 % respectively. Also, A isolate reflected highest hydrocarbons derivatives consumption comparing with C isolates which showed lowest hydrocarbons derivatives consumption %. Random amplified polymorphic DNA (RAPD) and SDS-PAGE techniques were employed to evaluate genetic similarity which led to explain molecular base for varied biodegradation activity for A, B and C isolates. Interestingly, A isolate which remarked with superior biodegrading activity reflected highest genetic polymorphism. On the other hand, highly genetic similarity was corresponded to decreasing degrading activity for isolate C. positive correlation was detected between genetic polymorphism and biodegrading capability for yeast isolate.
Keywords
Yeasts isolate
Hydrocarbon derivatives degradation
18S rRNA
Random amplified polymorphic DNA (RAPD)
SDS-PAGE techniques
1 Introduction
Hydrocarbons derivatives consider associated environmental pollutants with petroleum bioproduct which influence on human digestive. Compounds exposure causes a serious damage to liver and blood system which led to decreasing body responses and even death (Mitra and Roy, 2011). Thus initiate new biodegradation strategies would be essential for removing hazardous hydrocarbons from the ecosystem.
A wide variety of yeasts, bacteria and filamentous fungi are capable of consumption hydrocarbon slowly which oriented to accelerate these strategies due to its safety. Identification potentials microorganisms consider the main concern for develop biodegradation strategy.
Many studies cleared fungi superiority for hydrocarbon derivatives degradation over bacteria in contaminated soils (Prenafeta-Boldú et al., 2019) for different causes. Firstly, induction of several specific enzymes such as lignin peroxidase. Moreover, increasing growth rate in highly contaminated ecosystems. Finally, unique enhanced access to hydrocarbons through mycelia network (Bielèik et al., 2019). Thus, Penicillium and Fusarium, which were isolated from the polluted soil, were evaluated as bioaugmentation tool. Beside remarkable biodegradation cabability of total petroleum hydrocarbons (TPH), 5-ringed benzo(a)fluoranthene and benzo(a)pyrene were reduced. Also, 6-ringed benzoperylene decreased by a 28 % (Medaura et al., 2021).
Random Amplified Polymorphic DNA (RAPD)-PCR was employed as molecular fingerprinting to estimate genetic similarity among distinguishable yeast isolates with biodegradation activity.
Recently, SDS-Page) consider a powerful tool for monitoring and detecting variation among different microorganisms and has been used increasingly in microbial Systematic (Brarns et al., 1991; Hall et al., 1969; Johnston et al., 1988; Ochiai and Kawamoto, 1995).
This investigation was carried out to isolate and identify local petroleum hydrocarbon-degrading yeasts which characterized with biodegrading hydrocarbons. Moreover, 18S rRNA gene sequence was employed for yeast identification. Additionally, biodegradation derivatives were monitored to evaluate biodegrading activity for potentials yeast isolate. Finally, genetic similarity among isolates yeast with biodegradation abilities was evaluated through Random Amplified Polymorphic DNA (RAPD) molecular marker technique.
2 Materials and methods
2.1 Sediment sampling
Manual sampling device was used to collect sediments from three points of Ras Tanura City (26°38′38″N and 50°9′33″E), eastern province, Kingdom of Saudi Arabia which located on the Arabian Gulf surrounded by seawater. Samples were kept into sterile bags and directly transferred to laboratory in iced containers in less than 24 h to carry out enrichment step.
2.2 Yeast isolation
To enrich isolate yeast, Nutrient Broth (NB) (Oxoid, RM175.00) was used. After incubation for three days at 30˚C and agitation 200 rpm, Nutrient Agar (NA) (Oxoid, CM0003B) media was applied for culturing 10-3–10-5 sample dilutions.
2.3 Preliminary test for degradation ability
To evaluate biodegradation capability for yeast isolates, Natural Sea Water agar (NSWA) plates containing petroleum oil 0.5 % (v/v) was applied. After incubation at 30 °C for one week, nominated yeast isolate were selected for further studies.
2.4 Molecular identification of yeasts
18S rRNA was applied as molecular marker to identify yeast isolates. Total genomic DNA was purified via Yeast DNA Extraction Kit (Thermo Fisher, USA, 78870) according to manufacturer protocol. General eukaryotic primers EukA (5-AACCTGGTTGATCCTGCCAGT-3) and EukB (5-TGATCCTTCTGCAGGTTCACCTAC-3) primers were employed to amplify 18S rRNA subunits (Petroni et al., 2002). Amplification was carried out through Peqstar gradient thermal cycler (VWR, Germany) with following conditions, 95 °C for 5 min; 94 °C for 1 min, 52 °C for 1 min and 72 °C for 2 min (35 cycles); and 72 °C for 10 min (Hassanshahian et al., 2012). To visualized data, 1.5 % agarose gel electrophoresis (Ausubel et al., 1999) was used via Digi-doc it gel documentation sys-tem (UVP, UK). Amplicons were purified through Gene JET PCR Purification Kit (Thermo Scientific). ABI PRISM 3100 Genetic Analyzer was applied for sequenced and performed by Macrogen In. Seal, Korea. Aligned sequences were analyzed on NCBI website (https://www.ncbi.nlm.nih.gov/webcite) using BLAST to identify yeast isolate. Multiple sequence alignment, and molecular phylogeny was performed using ClusteralW software analysis (https://www.Clus-teralW.com). Nucleotide sequence were submitted to NCBI database to get the accession numbers.
2.5 Monitoring yeast growth and degradation ability
Colorimetric method was employed to evaluate growth rate with biodegradation activity. NSWA media supplemented with0.5 % crude petroleum oil (M1) was employed for yeast culturing with shaking (150 rpm) at 30 °C for 72 h. after adjusting media pH was to 7.0, microbial growth (OD600) was measured as optical density (OD) at a wave length of λ600 nm. Chloroform extraction method was used to measure oil consumption gravimetrically in acidified medium (APHA, 1998).
2.6 Hydrocarbons and some derivatives biodegradation potentials
Aromatic derivatives hydrocarbons (naphthylamine, phenol, naphthaleneethyldiamine, phenanthrene, naphthalene 2-sulfonate and naphthalene) and aliphatic (nhexane, n-heptane and n-pentadecane) was applied as marker for biodegradation cabbalism for yeast isolates. 600 mg/l was used for each hydrocarbon or derivative in NSWA medium. Two original yeast cultures were applied for inoculating NSW media supplemented with separate different aromatic hydrocarbons after grown yeast cultures for three days at 30 °C with agitation at 200 rpm. Spectrophotomoteric method was used for growth tracking. GC (Shimadzu GC-17A) and HPLC (Beckman system Gold 126 Solvent Module) were used for evaluating residual non-degraded hydrocarbons. Varied concentrations of naphthalene and phenol (250 to 5000 mg/l) were applied to determine degradation efficiency of the isolate.
2.7 RAPD analysis
Random amplified polymorphic DNA-PCR (RAPD-PCR) fingerprinting were studied via four random primers OPA-02 (5′-TGCCGCGCTG-3′), OPA-03 (5′-AGTCAGCCAC-3′), OPA-11(5′-CAATCGCCGT-3′) and OPA-15 (5′-TTCCGAACCC-3′) (Martorell et al., 2005). PCR master mix preparation, thermal cycler program and amplicons visualization were performed according to Martorell et al. (2005). Binary system was used to convert fragment profile to 0,1 matrix data with Digi-Doc it (gel documentation system, UVP, England) integrated with Totallab software analysis (Total.lab.v.1.1).
3 Results
Among nine yeast isolates from enrichment cultures, only three isolated reflected superior growth rate under crude oil application. 18S rRNA gene was amplified, eluted, sequence and allingemented and submitted to Sequence Database of the National Center for Biotechnical Information (NCBI) database. Phyllogenetic tree was constructed among three identified isolates and highest similarity isolates (Fig. 1.).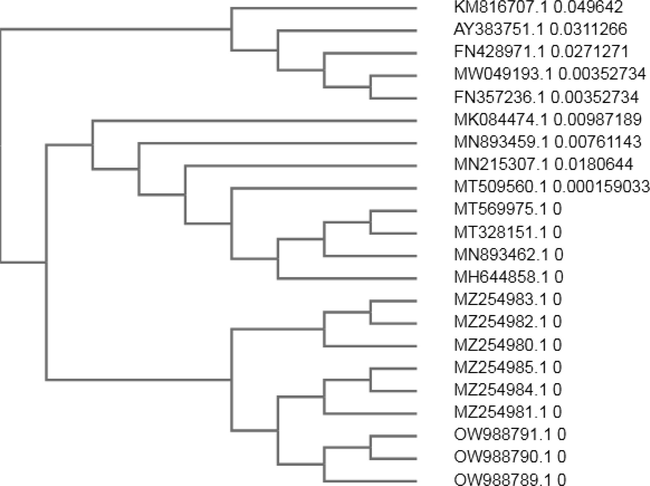
Constructed Phyllogenetic tree among three identified isolates and higest similarity isolates.
Partial sequence of 18Sr RNA isolates A, B and C has been submitted into GeneBank under accession numbers OL659287, OL659288 and OL659289. Based on similarity sequences data of 18Sr RNA multialignment sequence, isolates A, B and C were identical to Rhodotorula ingeniosa, Rhodotorula mucilaginosa and Candida parapsilosis with 100 % identity respectively (Table 1).
Isolates
Identifying
Accession No
A
Rhodotorula ingeniosa
OL659287
B
Rhodotorula mucilaginosa
OL659288
C
Candida parapsilosis
OL659289
Many yeast strains utilize aromatic hydrocarbons as sole carbon source (Xu et al., 2018). More light was added to our finding by Chandran and Das (2012). They identified and characterized five different yeast species as Cryptococcus laurentii, C. tropicalis, Rhodotorula mucilaginosa, Trichosporon asahii and Candida rugosa with remarkable biodegradation activity. Additionally, yeast isolate with biodegradation capability which identified as C. viswanthii was employed yeast isolate to biodegrade mixture of low and high molecular weight poly aromatic hydrocarbons (PAHs), gasoline, lubricating and vegetative oils from the contaminated soil (Hesham et al. (2009)).
3.1 Biodegradation assays
As shown by Table 2, three yeast isolates showed varied potentials for oil consumption. A isolate reflected higest oil degradation (8.2 %) comparing with lowest degradation % (5.5 %) which remarked C isolate. Positive correlation was detected between Oil consumption % and growth rate. Isolate B revealed moderate biodegradation activity comparing with A and C isolates. Positive relation was detected between degradation ability and growth rate.
Yeast isolates
OD600
Oil consumption %
A
6.22
8.2
B
2.78
6.4
C
0.48
5.5
Many studies support our findings for employed yeast as oil biodegradation microorganism by Kazemzadeh et al., 2022; Oyewole et al., 2022).
More light was added by Hesham et al., (2006) for using enrichment technique to select potential yeast isolates. They cleared that enrichment technique consider a powerful tool for tracking promising oil biodegrading ability of yeast strains. Furthermore, different researches selected and identified hydrocarbon biodegrading yeasts from varied ecosystems (Manee et al., 1998; Zinjarde and Pant, 2002). In accordance with our findings for isolating yeast from water, hydrocarbon degrader yeast strains was identified and characterized as C. albicans and Candida maltose in lagoon water from Nigeria (Matthew et al. (2008). Our results are in agreements to those of Zinjarde et al. (1997); they identified Y. lipolytica from oil-polluted Indian sea water.
3.2 Hydrocarbons and some derivatives degradation potentials of C.tropicalis strain A
Three strains capability for using varied hydrocarbons and some derivatives at 600 mg/L level was assayed after incubation for 72 h in NSW medium supplemented with selected compounds as a sole carbon source.
As shown by Table 3 and Fig. 2, biodegradation ability for hydrocarbons derivatives was varied significantly. Yeast strain C was superior for utilizing naphthylamine, phenol, naphthyl ethyl diamine and phenanthrene. Furthermore, negative correlation was detected between growth rate and length of aliphatic chain. Generally, remarked n-pentadecane with expanded chain length caused decreasing growth rate for three isolates.
Isolates
hydrocarbons derivatives
n-pentadecane
phenol
Naphthyl ethyl diamine
naphthylamine
OD600
consumption %
OD600
consumption %
OD600
consumption %
OD600
consumption %
A
0.255
0.74
1.257
71.25
0.335
36.84
0.474
36.84
B
0.188
0.55
0.955
66.31
0.247
23.54
0.322
23.54
C
0.039
0.000
0.491
53.60
0.207
20.00
0.279
20.00
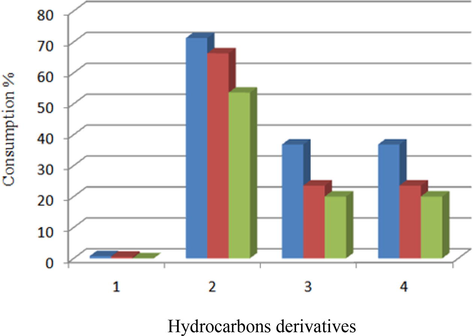
Consumption % of hydrocarbons derivatives for three yeast isolates.
Primers
Isolates
A
B
C
Total fragments
Polymorphic fragments
Polymorphism %
Total fragments
Polymorphic fragments
Polymorphism %
Total fragments
Polymorphic fragments
Polymorphism %
1
15
10
66.7
13
7
53.8
13
7
53.8
2
13
8
61.5
14
8
57.1
14
5
35.7
3
16
10
62.5
17
8
47.1
16
2
12.5
4
15
11
73.3
17
6
35.3
14
5
35.7
5
16
9
56.3
12
7
58.3
14
6
42.9
Average of Polymorphism %
64.1
50.3
36.1
Genetic Stability
%35.9
49.7
63.9
Interestingly, isolate A did not show any biodegradation activity against n-pentadecane. Lowest biodegrading rate was characterized n-pentadecane with 0.55 and 0.74 of consumption % for A and B respectively. By contrary, three isolates (A, B and C) reflected highly phenol consumption rates (53.60, 66.31 and 71.25 %) respectively. Based on previous data, n-pentadecane and phenol degradation abilities for isolate C were estimated via supplemented NSW medium with different concentrations ranging from 200 to 4000 mg/L. As shown by Table 3, negative correlation was detected between isolate C growth rate and naphthalene concentrations. Highly naphthalene removing was detected at 200 mg/l. on the other hand, increasing naphthalene concentration at 4000 mg/l due to dramatic decrease of naphthalene removal (25 %). Highly phenol removal (52 %) was recorded at moderate concentration (1200 mg/L).
Our obtaining results for varied biodegradation abilities of yeast against hydrocarbons derivatives was in agreements with Kottb et al. (2019). They explained decreasing biodegrading naphthylamine and naphthalene 2- sulphonate cause of their lethal effect. More support was added to our results for increasing yeast biodegrading ability after incubation by (Lopes et al., 2019). They cleared that transferred oxygen mass was increased which due to an improvement on cellular growth and reducing sugars consumption. Also, increasing special receptor sites for binding hydrocarbons which uptake hydrocarbon uptake and may be play a key role for transfer hydrocarbons into the cell. Furthermore, catabolic pathway was accelerated via energy which generate via enzymes that introduce oxygen into the carbon.
To evaluate genetic similarity among three yeast biodegrading strains (A, B and C). Random amplified polymorphic DNA-PCR (RAPD-PCR) fingerprinting with five primers was applied. As shown by Fig. 3 and table, all yeast strains showed different amplified fragments and polymorphism %. Generally, isolate A was superior for total amplified, polymorphic fragments and decreasing of genetic stability % comparing with isolates B and C. A yeast isolate reflected 75 fragments as total fragments with 64.1 % as polymorphism %. 73 amplicons with 50.3 % as polymorphism % were characterized B isolate. Lowest amplicons (71) and polymorphism % (36.1 %) were recorded for C isolate. A yeast isolate showed lowest genetic stability % (35.9 %) comparing with B and C isolates which reflected moderate and high polymorphism % (49.7 and 63.9) respectively. Distinguishable decreasing of genetic stability for isolate A may be considering the key role for biodegrading superiority. Based on RAPD data phyllogenetic tree was constructed. As a result for remarked similarity, isolates B and C were located in the same cluster with 55 % of genetic similarity. A isolate was represented as separate cluster due to lacking of genetic stability %. In accordance with our results for applied Random amplified polymorphic DNA-PCR as fingerprinting technique to understand variation of biodegradation activity among three yeast isolates, Hesham et al., (2018) used RAPD and proved correlation between random amplified polymorphic DNA profile and the geographic origin of seven toluene-degrading red yeasts (A5, A6, A12, A20, A26, A29, and A38). More support was added to our RAPD results by Meroth et al., (2003) and EL-Fikyet al., (2012). They indicated that RAPD-PCR technique consider a powerful useful tool for yeast identification and genetic diversity evaluation.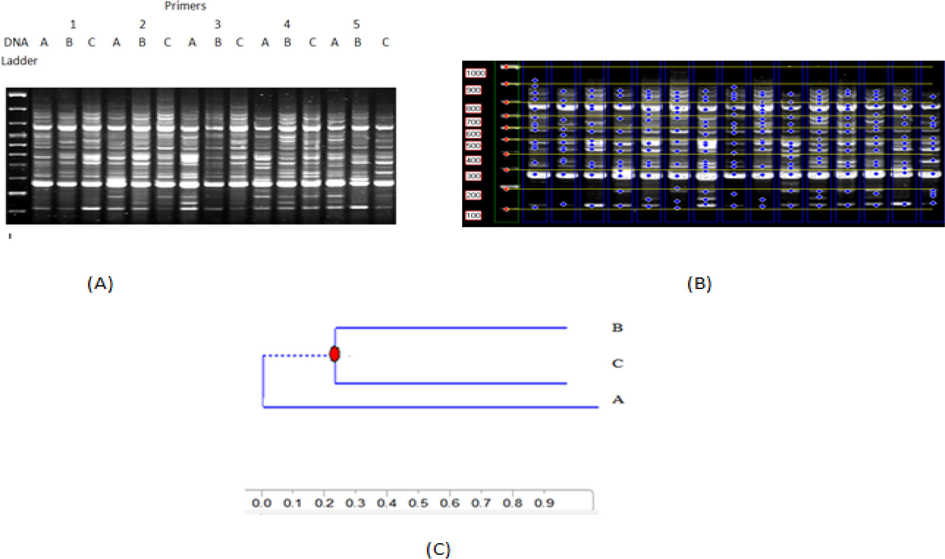
Random Amplified Polymorphic DNA (RAPD) fingerprinting results for five random primers for A, B and C yeast isolates.
3.3 Protein profile study
Investigating protein profile for three yeast isolates consider a powerful tool to clear molecular base for genetic diversity among yeast isolates with varied biodegrading ability. Different polymorphism % was detected for three isolates. As shown by Fig. 4 and Table 5, isolate A reflected highest polymorphism % (69.2 %). On the other hand, lowest protein polymorphism % was recorded for isolates C (20 %). Isolate B showed moderate polymorphism % (37.5) based on protein profile data. Designed phyllogenetic tree based on protein profile indicated superior diversity for isolate A which located separately at 8 % of genetic similarity. Unique protein profile for isolate A could be led to its distinguishable activity for oil biodegradation.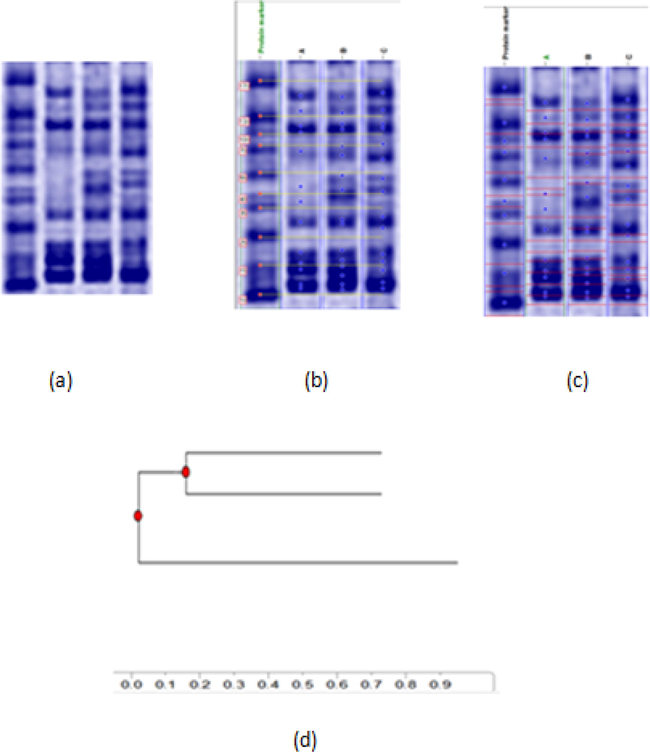
Protein fingerprinting profile (a), band detection (b) and molecular weight calculation (c) and phyllogentic tree (d) for A, B and C yeast isolates.
Yeast isolates
A
B
C
Total bands
13
16
15
Polymorphic bands
9
6
3
Polymorphism %
69.2
37.5
20.0
Our findings for using SDS-PAGE to explain biodegradation activity based on protein profile diversity was supported by Hofling et al. (2018). They used gel electrophoresis to investigate yeast soluble proteins of detect biodegradable plastic-degrading enzyme (PaE) from Pseudozyma antarctica JCM 10317. Corresponding patterns showed unique 36 polypeptides ranged from 14.4 to 200 kDa. Different nutritional compositions could explain protein patterns diversity as a result to express varied proteins derived from alternatives metabolic pathways. More light was added to our findings by Yu et al., (2015). They applied SDS-PAGE and N-terminal sequencing to understand Polyacrylamide (PAM) degrading by Pseudomonas putida which indicated conversion PAM amide groups into carboxyl groups by a PAM-induced extracellular enzyme from the aliphatic amidase family.
4 Conclusion
Three yeasts which isolated from contaminated soil of Khafji oil fie from of Ras Tanura City (Eastern Province, Saudi Arabia) were assayed for hydrocarbon biodegradation capability. Through 18S rRNA gene sequence which employed as molecular identification tool, three yeast isolates identified and submitted under accession numbers OL659287, OL659288 and OL659289 and identical as Rhodotorula ingeniosa, Rhodotorula mucilaginosa and Candida parapsilosis with 100 % identity respectively. A, B and C isolates reflected 8.2, 6.4 and 5.5 % of biodegradation % respectively. According to Random amplified polymorphic DNA-PCR (RAPD-PCR) fingerprinting results, three isolates reflected varied Genetic Stability (35.9, 49.7 and 63.9 %, respectively) which could be explain the differences for biodegradation activity.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Ausubel, F.M., Brent, R.,Kingston, Moore, R. E., Seidman, J.G., Smith, J. A., Struhl, K., 1999. assimilate hydrocarbons. Anto. Leeuwen., 34, 441– 457. (5), 394–398.253.
- The role of active movement in fungal ecology and community assembly. Mov. Ecol.. 2019;7:36.
- [CrossRef] [Google Scholar]
- Chandran, P., Das, N., 2012. Role of plasmid in diesel oil degradation by yeast species isolated Current protocols in molecular biology, John Wiley and Sons Inc, New York, NY. degrading four and five ring aromatic hydrocarbons, Ann. Microbiol. (56), 109—112 18.Dent. Res., 71: 961.
- Quality parameters and RAPD—PCR Differentiation of Commercial Baker's Yeast and Hybrid Strains. J. Food Sci.. 2012;77(6):2012.
- [Google Scholar]
- Approach to taxonomy of Phytophthora through acrylamide gel-electrophoresis of protein. Phytopathology.. 1969;59:770-774.
- [Google Scholar]
- Isolation and characterization of two crude oil-degrading yeast strains, Yarrowia lipolytica PG-20 and PG-32, from the Persian Gulf. Mar. Pollut. Bull.. 2012;64(7):1386-1391. Epub 2012 May 22 PMID: 22622152
- [CrossRef] [Google Scholar]
- Isolation and molecular genetic characterization of a yeast strain able to degrade petroleum polycyclic aromatic hydrocarbons. Afr. J. Biotechnol.. 2009;8(10):2218-2223.
- [Google Scholar]
- Isolation and identification of a yeast strain capable of degrading four and five ring aromatic hydrocarbons. Ann. Microbiol.. 2006;56:109.
- [CrossRef] [Google Scholar]
- Degradation of Toluene Hydrocarbon by Isolated Yeast Strains: Molecular Genetic Approaches for Identification and Characterization. Russ. J. Genet.. 2018;54:933-943.
- [CrossRef] [Google Scholar]
- Differentiaion and numerical analysis of oral yeast based on SDS-PAGE profiles infleumce of the culture media on the whole-cell protein extracts. Brazil. J. Biol., 61(3), 507–516. Hydrocarbon gel electrophoresis. Yeast. 2018;4:191-198.
- [Google Scholar]
- Karyotyping of yeast strains of several genera by field inversion gel electrophoresis. Yeast.. 1988;4(3):191-198. PMID: 3201829
- [CrossRef] [Google Scholar]
- Kazemzadeh omayeh, Zarrindokht Emami-Karvani, Nafiseh Sadat Naghavi, Giti Emtiazi, 2022. Production of surface-active sophorolipid biosurfactantand crude oil degradability by novelRhodotorulamucilaginosastrain SKF2. J Surfact Deterg.2022;1–16.
- Molecular monitoring of spoilage yeasts during the production of candied fruit nougats to determine food contamination sources. Int. J. Food Microbiol.. 2005;101(3):293-302. PMID: 15925712
- [CrossRef] [Google Scholar]
- Isolation and characterization of hydrocarbon-degrading and biosurfactant-producing yeast strains obtained from a polluted lagoon water. World J. Microbiol. Biotechnol.. 2008;24:2539-2545.
- [Google Scholar]
- Bioaugmentation of Native Fungi, an Efficient Strategy for the Bioremediation of an Aged Industrially Polluted Soil With Heavy Hydrocarbons. Front by Isolated Yeast Strains: Molecular Genetic Approaches A. for Identification and Characterization. Russ. J. Genet.. 2021;54(8):933-943.
- [Google Scholar]
- Identification and population dynamics of yeasts in sourdough fermentation processes by PCR-denaturing gradient gel electrophoresis. Appl. Environ. Microbiol.. 2003;69(12):7453-7461. PMID: 14660398; PMCID: PMC309968
- [CrossRef] [Google Scholar]
- Mitra, S. and Roy, P., 2011. BTEX: a serious ground-water contaminant, Res. J. Environ. Sci, novel approach to bacterial taxonomy: application to the genus Arthrobacter. Biosci. Biotechnol. Biochem. (62), 1679-1687.19.
- Two-dimensional gel electrophoresis of ribosomal proteins as a novel approach to bacterial taxonomy: application to the genus Arthrobacter. Biosci. Biotechnol. Biochem.. 1995;59(9):1679-1687. PMID: 8520111
- [CrossRef] [Google Scholar]
- Oluwafemi A Oyewole, Clement S Olusanya, Olusegun I Akinlade, Yusuf O Abdulwasiu, Anthony O Kelechi and Michael E Okoro Biodegradation of selected petroleum hydrocarbons using indigenous microorganisms. PG-32, from the Persian Gulf. Marine pollution bulletin. (64), 1386-91.
- Prenafeta-Boldú, F. X., de Hoog, G. S., and Summerbell, R. C., 2019. “Fungal Communities in Hydrocarbon Degradation,” in: T. McGenity (ed.) Microbial Communities Utilizing Hydrocarbons and Lipids: Members, Metagenomics and Ecophysiology. Handbook of Hydrocarbon and Lipid Microbiology, (Cham: Springer), doi: 10.1007/978-3-319-60063-58-2.
- Petroleum Hydrocarbon-Degrading Bacteria for the Remediation of Oil Pollution Under Aerobic Conditions: A Perspective Analysis. Front. Microbiol.. 2018;9:2885.
- [Google Scholar]
- Hydrocarbon degraders from tropical marine environments. Int. J. Food Microbiol.. 2002;101:293-302.
- [Google Scholar]







