Assessment of the relationships between IL-17A polymorphisms and the risk to acute lymphoblastic leukemia in Saudi population
⁎Corresponding authors. lmansour@ksu.edu.sa (Lamjed Mansour)
-
Received: ,
Accepted: ,
This article was originally published by Elsevier and was migrated to Scientific Scholar after the change of Publisher.
Peer review under responsibility of King Saud University.
Abstract
The most studied IL-17 family member is IL-17A. It has been implicated in the pathogenesis of many disorders, including inflammatory, autoimmune and cancer disorders. However, the role of this cytokine in the pathophysiology of acute lymphoblastic leukemia (ALL) remains poorly studied. The purpose of this study was to investigate the role of IL-17A gene single-nucleotide polymorphisms in the occurrence of ALL in Saudi population. The current study included 136 patients with ALL and 114 age and gender-matched healthy controls. The relationship between three functional SNPs of IL-17A gene; rs3748067, rs3819025 and rs8193036 was assessed in both ALL and healthy control following five inheritance models. In addition. the level of IL-17A mRNA was analyzed by quantitative polymerase chain reaction (qRT-PCR). The results revealed a statistically significant protective effect of the rs3748067 CT genotype against ALL in dominant, overdominant and additive models (p < 0.05). No association was observed between the rs3819025, and rs8193036 and the risk to develop ALL (P > 0.05). Haplotype analysis showed association of CTG (OR = 0.52, 95 %CI 0.29–0.94, p = 0.03), and TCG (OR = 0.45, 95 %CI 0.25–0.81, p = 0.009) of rs3748067, rs8193036 and rs3819025 with higher risk of ALL. Our results showed that IL-17A promoter rs3748067 variant may reduces the risk to ALL.
Keywords
IL-17
Single nucleotide polymorphism
Acute lymphoblastic leukemia
Saudi Arabia
1 Introduction
Childhood acute lymphoblastic leukemia (ALL) is the most common malignancy diagnosed in children, characterized by the proliferation of lymphoid progenitor cells in the bone marrow, blood, and extramedullary sites. Ninety percent of ALL cases occur in children, whereas it is a devastating disease when it occurs in adults (Inaba et al., 2013). In most cases, relapses, treatment resistance, and serious infections are the leading causes of death. Tissue microenvironmental pro-survival signals, such as intricate crosstalk between ALL cells and CD4+ T cells, mesenchymal stromal cells, and various cytokines, help to maintain leukemic clones and promote chemotherapy resistance in adult ALL patients (Purizaca et al., 2012).
Th17 cells, were determined as unique subset of T helper cells that play an important role in both innate and adaptive immunity. They secretes a high quantity of interleukin (IL)-17, known as pro-inflammatory cytokine (Harrington et al., 2005; Iwakura et al., 2011). IL-17 includes six family members (IL17A to F) which bind to five receptors (IL-17RA to E) (Kawaguchi et al., 2004). IL-17A is one of the most important cytokines of IL-17 family members and usually known as IL-17. It plays an important role in autoimmune diseases, chronic inflammatory diseases and malignancies (Dai et al., 2016a; Su et al., 2010). Corroborating studies have reported that IL-17A can induce the expression of various inflammatory chemokines, cytokines and adhesion molecules involved in inflammation-related cancer (Dai et al, 2016b; Song & Qian, 2013). It has been reported to play regulatory role in cellular proliferation, angiogenesis and metastasis. In addition, it may contributes to the increased susceptibility to tumors (Yang et al., 2014). In fact, IL-17 production is tightly regulated, and its over-production may induce inflammation, that were associated with a myriad of pathologies, including allergies, celiac disease, and cancers (Bedoui et al., 2018; Cui et al., 2012; Lee et al., 2018; Numasaki et al., 2003; Tseng et al., 2014; Veldhoen, 2017). Also, It has been reported that IL-17A is overexpressed in a variety of cancers, including ovarian, gastric, prostate, colorectal, lung, hepatocellular, and esophageal(Kirshberg et al., 2011; Kuang et al., 2010; Le Gouvello et al., 2008; Lv et al., 2011; Miyahara et al., 2008; Sfanos et al., 2008; Zhang et al., 2008). Single nucleotide polymorphisms (SNPs) are as the most common type of genetic diversity in the human genome usually associated with a detrimental phenotype (Erichsen & Chanock, 2004; Li et al., 2018). They can be found in different parts of the genes including the promoters, exons, introns, and also the 5′- and 3′- UTRs (Deng et al., 2017). This can results in the production of an altered protein that could cause functional impairment of the protein or influence its expression implications, especially if they are located in the coding regions or regulatory regions of the genes (Fernández et al., 2016). In this context, large number of SNPs, especially those affecting the function of some molecules reported to play important role in tumorigenesis, have been investigated and were found to be associated with the risk to develop cancer disease or to influence the outcome of pharmacological treatments (Deng et al., 2017; Erichsen & Chanock, 2004; Lange et al., 2021). Currently, many studies have reported the association of some SNPs in the IL-17 gene with cancer diseases including gastric (Elshazli et al., 2018), breast (Wang et al., 2012), colorectal (Li et al., 2022) and thyroid cancer (Bertol et al., 2022). At the present, there is still a lack of data on the relationship between IL and 17A gene polymorphism and disease susceptibility to ALL. In Saudi Arabia, few studies have been performed on the association between some polymorphisms in the immune molecules such as TLR-4 and HLA-G with ALL, and strong relationships were found between some poliorphism and the occurrence of the disease a (Al-Tamimi et al., 2022; AlKhulaifi et al., 2022).
Therefore, in this study we genotyped three representative functional IL-17A SNPs; rs3748067(C>T), IL-17A rs3819025 (G>A), and IL-17A rs8193036 (C>T) polymorphisms and examined their possible association with ALL in a case/control study in Saudi population.
2 Material and methods
2.1 Patients and healthy control
The present study has enrolled 136 Saudi patients, including 20 (15%) females and 115 (85%) males) with acute lymphoblastic leukemia (ALL) living in Riyadh city. Control group is composed of 114 healthy volunteers including 37 female (32.5%) and 77 male (67.5%) age- and sex-matched. The mean age of the study population was 22.45 ± 20.27 years for the patients with ALL and 18.68 ± 15.53 years for the healthy controls (Table 1). None of control subjects reported personal or family history with ALL and any other chronic or immune diseases.
| Characteristics | ALL (136) | Control (114) | |
|---|---|---|---|
| Male | 116 | 77 | |
| Gender (number) | Female | 20 | 37 |
| Age (average ± SD) | 22.45 ± 20.27 | 18.68 ± 15.53 | |
2.2 Samples collection and DNA extraction
3 ml of blood samples were drawn aseptically from each study groups and stored at −20°C in ethylenediaminetetraacetic acid (EDTA)-containing tubes before analysis. The DNA of peripheral blood from ALL patients and healthy controls was extracted by QIAamp DNA Mini Kit (Qiagen, Germany) according to the manufacturer’s instructions. The concentration of DNA was measured with a Nanodrop ND-2000c spectrophotometer (Thermo Scientific, Wilmington, DE, USA).
2.3 Single nucleotide polymorphisms (SNPs) selection
In this research, we identified three single nucleotide polymorphisms (SNPs) in IL-17A gene (rs3748067, rs3819025 and rs8193036) through the dbSNP databases (https://www.ncbi.nlm.nih.gov/snp/)). SNP were selected according to their minor allele frequency (MAF) ≥5%.; Hardy-Weinberg equilibrium (HWE) P value cut-off > 0.005 (Table 2).
| SNP ID/assay ID | Chromosome position | Nucleotide change |
Region | MAF imHuman populations (1000genomes Study) | Present study | ||||
|---|---|---|---|---|---|---|---|---|---|
| Global | European | South Asian | East Asian | Qatari | |||||
| rs3748067 | Chr6/ 52,190,541 | C/T | 3-’UTR | T = 0.09 | T = 0.10 | T = 0.09 | T = 0.09 | T = 0.060 |
T = 0.11 |
| rs3819025 | Chr 6/ 52,186,476 | G/A | Intron Variant | A = 0.07 | A = 0.06 | A = 0.16 | A = 0.22 | A = 0.09 | A = 0.16 |
| rs8193036 | Chr 6/ 52,185,695 | C/T | Upstream Variant | T = 0.74 | T = 0.75 | T = 0.56 | T = 0.29 | T = 0.19 | T = 0.16 |
MAF: minor allele frequency.
IL-17 genotyping of the three SNPs was performed by vIC- and FAM-labelled allelic discrimination method, using assay-on-demand TaqMan assays ordered from Applied Biosystems according to the manufacturer’s instructions using an ABI Prism 7500 Real-time PCR system (Applied Biosystems, Foster City, USA). Real-time PCR was implemented in 10 μl a reaction system containing 0.26µl 2x SNP Genotyping Assay, 5.5µl 2xPower Taq MasterMix Mix, 2.24µl Nuclease-Free Water, and 2 µl DNA template (100 ng/μl).
The PCR conditions for IL-17A rs3748067, rs3819025 and rs8193036 were 1 cycle at 95°C for 10 min followed by 40 cycles (95°C for 15 s, 55°C for 30 s, and 72°C for 30 s) and a final extension at 72°C for 5 min. For confirmation, about 5% of the samples were randomly chosen for repeat genotyping.
2.4 Quantitative real-time PCR analysis
Briefly, total RNA was extracted using PureLink™ RNA Mini Kit (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. Roughly, 1 μg of total RNA from every sample was used to synthesize cDNA with PrimeScript RT reagent Kit (Takara Bio Inc., Dalian, China). The reverse transcription reaction was performed at 37 °C for 15 min, followed by 5 s at 85 °C.
Real-time quantitative PCR was performed using an ABI Prism 7500 Real-time PCR system (Applied Biosystems, Foster City, CA, USA) in accordance to the manufacturer’s protocols. The real-time PCR was performed as reported by (AlKhulaifi et al., 2022). Briefly, in a final volume of 10 μl, 5 μl of SYBR Green Real-time PCR Master Mix, 3.2 μl of DEPC- treated H2O, 1 μl of cDNA, 0.8 μl of the forward and reverse primers and 1 μl of cDNA. For specific 17A amplification, the sequences of primers used were F: 5′-CCGTTTTATCACGGAGGTGGT-3′and R: 5′-CTGCCTAAATGCCTCAGGGG −3′. ′.All experiments were carried out in triplicate. Target results were expressed relative to the number of GAPDH transcripts used as an internal control. The primer sequences of GAPDH were: F: 5′-TCTCCTCTGACTTCAACAGCGAC-3′ and R: 5′-CCCTGTTGCTGTAGCCAAATTC-3. For the relative quantification, data were analyzed by the comparative 2−ΔΔCt method and normalized to the average of housekeeping gene GAPDH.
2.5 Statistical analysis:
Association study including the model with the lowest Akaike information criteria (AIC) value was considered as the best one. Odds ratios (OR) with 95 % confidence intervals (CI) were calculated using logistic regression.
All SNPs were tested for deviation to Hardy–Weinberg equilibrium using the Chi-square test. The relative risk associated with alleles and genotypes was calculated as an odds ratio (OR) with a 95% confidence interval (CI) for five inheritance models including co-dominant, dominant, recessive, over-dominant and log-additive models were performed using the web-based SNPStats software program (Solé et al., 2006). Analysis of the linkage disequilibrium (D′) between the IL-17A SNPs was also estimated using the SNPStats and Haploview. softwares.
3 Results
A total of 136 eligible patients and 114 controls were genotyped for the three target SNPs of IL-17A (rs3748067 (C>T), rs3819025 (G>A) and rs8193036 (C>T)).
All patients and controls have been tested for IL-17A rs3748067, rs3819025, and rs8193036) polymorphisms, using TaqMan methods.
The genotype distribution of IL-17A rs3748067(C>T) and rs8193036 (C>T) polymorphisms in the control cohorts followed Hardy–Weinberg equilibrium, except for IL-17A rs3819025 polymorphism (G/A).
3.1 Relationships between IL and 17A SNP and risk to ALL
Table 3 summarizes the allelic and genotypic distributions of rs8193036 (C>T), rs3819025 (G>A) and rs3748067(C>T) polymorphisms among ALL patients and healthy controls and the strength of association with disease following logistic regression. For the rs3748067, protective effects were observed for the CT genotype in codominant model; CT vs CC (OR = 0.47, 95%CI (0.25–0.91) and p = 0.024) for overdominant model; CT versus CC + TT (OR = 0.50, 95%CI (0.26–0.91) and p = 0.024), for dominant model; CT + TT vs CC (OR = 0.48, 95%CI (0.26–0.87) and p= 0.015) and Log additive model (OR = 0.59, 95%CI (0.37–0.95) and p= 0.025),. Also, the T allele occurs with higher frequency in control (18%) compared to ALL patients (11%), suggesting a protective effect (OR = 0.53 95%CI (0.32–0.95) and p= 0.033) against ALL. Analysis of the rs8193036 C>T, shows higher frequency of T allele among healthy (23%) compared to ALL (16%), suggesting a protective effect (OR = 0.63, 95%CI (0.46–0.99) and p = 0.046). A higher frequency of TT and CT genotypes in healthy individuals, but the differences are not significant (p<0.05). For the rs3819025G>A, no significant association was found with ALL instead of some difference in the distribution (p<0.05).
| Locus | Model | Genotype |
ALL (%) N = 136 |
Controls (%) N = 114 |
OR (95 % CI) | P-value | AIC |
|---|---|---|---|---|---|---|---|
| rs8193036 C > T |
Alleles | C | 0.84 | 0.77 | 1 | ||
| T | 0.16 | 0.23 | 0.63(0.46 – 0.99) | 0.046 | |||
| Codominant | CC | 98 (72.1 %) | 71 (62.3 %) | 1.00 | |||
| CT | 33 (24.3 %) | 34 (29.8 %) | 0.70 (0.40–1.24) | 0.22 | 347.1 | ||
| TT | 5 (3.7 %) | 9 (7.9 %) | 0.40 (0.13–1.25) | 0.10 | |||
| Dominant | CC | 98 (72.1 %) | 71 (62.3 %) | 1.00 | |||
| CT + TT | 38 (27.9 %) | 43 (37.7 %) | 0.64 (0.38–1.09) | 0.1 | 345.9 | ||
| Recessive | CC + CT | 131 (96.3 %) | 105 (92.1 %) | 1.00 | |||
| TT | 5 (3.7 %) | 9 (7.9 %) | 0.45 (0.14–1.37) | 0.15 | 346.5 | ||
| Overdominant | CC + TT | 103 (75.7 %) | 80 (70.2 %) | 1.00 | |||
| CT | 33 (24.3 %) | 34 (29.8 %) | 0.75 (0.43–1.32) | 0.32 | 347.7 | ||
| Log-Additive | --- | --- | 0.67 (0.44–1.02) | 0.062 | 345.1 | ||
| HWE (P value) | 0.30 | 0.10 | |||||
| rs3819025 G > A |
Alleles | G | 0.82 | 0.84 | Ref | – | |
| A | 0.18 | 0.16 | 0.83 (0.5203–1.3319) | 0.5 | |||
| Codominant | GG | 93 (68.4 %) | 86 (75.4 %) | 1.00 | 1 | ||
| GA | 36 (26.5 %) | 20 (17.5 %) | 1.66 (0.90–3.10) | 0.10 | 347.1 | ||
| AA | 7 (5.2 %) 8 | 8 (7 %) | 0.81 (0.28–2.33) | 0.69 | |||
| Dominant | GG | 93 (68.4 %) | 86 (75.4 %) | 1.00 | |||
| GA + AA | 43 (31.6 %) | 28 (24.6 %) | 1.42 (0.81–2.33) | 0.22 | 347.1 | ||
| Recessive | GA + GG | 129 (94.8 %) | 106 (93 %) | 1.00 | |||
| GG | 7 (5.2 %) | 8 (7 %) | 0.72 (0.25–2.05) | 0.54 | 348.3 | ||
| Overdominant | GG-AA | 100 (73.5 %) | 94 (82.5 %) | 1.00 | |||
| GA | 36 (26.5 %) | 20 (17.5 %) | 1.69 (0.91–3.13) | 0.09 | 345.8 | ||
| Log-Additive | 1.16 (0.76–1.78) | 0.49 | 348.2 | ||||
| HWE (P value) | 0.187 | 0.0011 | |||||
| rs3748067 C > T |
Alleles | C | 0.89 | 0.82 | 1 | ||
| T | 0.11 | 0.18 | 0.53 (0.32–0.88) | 0.013 | |||
| Codominant | CC | 112 (82.3 %) | 79 (69.3 %) | 1.00 | |||
| CT | 19 (14 %) | 28 (24.6 %) | 0.47 (0.25–0.91) | 0.024 | 344.8 | ||
| TT | 5 (3.7 %) | 7 (6.1 %) | 0.50 (0.15–1.64) | 0.25 | |||
| Dominant | CC | 112 (82.3 %) | 79 (69.3 %) | 1.00 | |||
| CT + TT | 24 (17.6 %) | 35 (30.7 %) | 0.48 (0.26–0.87) | 0.015 | 342.8 | ||
| Recessive | CT + CC | 131 (96.3 %) | 107 (93.9 %) | 1.00 | |||
| TT | 5 (3.7 %) 7 | 7 (6.1 %) | 0.58 (0.18–1.89) | 0.36 | 347.8 | ||
| Overdominant | CC + TT | 117 (86 %) | 86 (75.4 %) | 1.00 | |||
| CT | 19 (14 %) | 28 (24.6 %) | 0.50 (0.26–0.95) | 0.033 | 344.1 | ||
| Log-Additive | 0.59 (0.37–0.95) | 0.025 | 343.6 | ||||
| HWE (P value) | 0.008 | 0.052 |
ALL acute lymphoblastic leukemia, OR odds ratio, 95 % CI 95 % confidence interval, *p < 0.05 was considered significant and are depicted in bold.
3.2 Haplotype analyses of IL-17A gene polymorphisms and risk to ALL
Table 4 shows the haplotype distributions of the IL-17A SNPs rs3748067(CT), rs8193036 (CT) and s3819025 (GA) in ALL patients and healthy control subjects. Among the 8 possible haplotypes six with ≥ 1% and representing cumulative of 99.25% were included in the analysis. The C-G-C haplotype was the most frequent among both healthy (51%) and patients (63%) and was considered as reference haplotype. Comparative analysis for the other 5 haplotypes shows protective effects of T-G-C (OR = 0.52, 95%CI (0.29–0.94) and p = 0.03) and C-G-T haplotype (OR = 0.45, 95%CI (0.25–0.81) and p = 0.009).
| rs8193036 | rs3819025. | rs3748067 | ALL | Control | OR (95 % CI) | P-value |
|---|---|---|---|---|---|---|
| C | G | C | 0.63 | 0.51 | Ref | |
| T | G | C | 0.09 | 0.16 | 0.52 (0.29–0.94) | 0.025 |
| C | G | T | 0.08 | 0.16 | 0.45 (0.25–0.81) | 0.0089 |
| C | A | C | 0.12 | 0.11 | 0.88 (0.48–1.61) | 0.74 |
| T | A | C | 0.05 | 0.04 | 0.85 (0.32–2.24) | 0.71 |
| T | G | T | 0.014 | 0.02 | 0.57 (0.13–2.53) | 0.46 |
Linkage disequilibrium (LD) analysis in control samples revealed LD between rs3748067 and rs3819025 (D′ = 0.67, p = 0.0042) (Fig. 1).
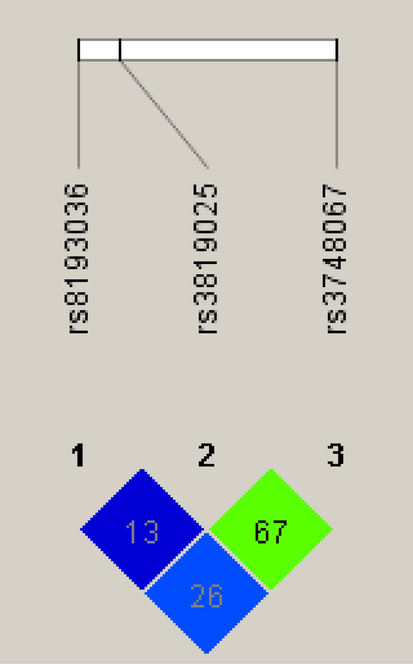
- Linkage disequilibrium (LD) plots of three SNPs in the IL-17A gene region. The plot was generated by Haploview. Numbers in squares show D' values between the SNPs expressed as percentages within the respective squares.
3.3 Relative mRNA expression of IL-17A
The expression of IL17A gene in patients was assessed by qRT-PCR. Analysis of the relative expression of mRNA level for ALL and healthy control subjects shows slightly higher level in control than in patients but the difference between groups is not significant (p < 0.05) (Fig. 2).
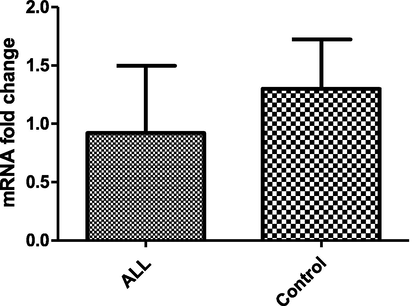
- The relativeIL-17A gene expression estimated using 2 − ΔΔCt method and normalized to the average of the GAPDH housekeeping gene. (mean ± SD).
4 Discussion
Th17 cells and their secretory cytokines are thought to play important role in autoimmune and inflammatory disorders but also a key cytokine in host protective immunity to infection (Agak et al., 2021; Hirota et al., 2011; Mease et al., 2015; Mills, 2022). Many studies have suggested that Th17 cells may have an effect on solid tumors, but their role in hematological malignancies is still being explored (Jin et al., 2019; Zhao et al., 2020). As well as, they have been shown to promote tumor cell survival, angiogenesis, and metastasis, through the activation of inflammatory signaling(Schetter et al., 2009; Wang et al., 2012; Wang et al., 2020; Wu et al., 2016; Yang et al., 2014).
Previous studies has linked IL17A polymorphisms to an increased risk to several types of cancer, including hepatocellular carcinoma, gastric cancer, acute myeloid leukemia, colorectal cancer, and papillary thyroid cancer (Dai , et al., 2016b; ELBassuoni et al., 2015; Hou et al., 2015; Lee et al., 2015; Li et al., 2022; Ma et al., 2015; Niu et al., 2017; Omrane et al., 2014; Omrane et al., 2015; Yang et al., 2014).
In the current study, we conducted a case/control study to evaluate the role of three selected SNPs of the IL-17A gene in the development of acute lymphoblastic leukemia (ALL) in Saudi population. Our analysis showed protective effects of the IL-17A rs3748067 polymorphism (located in the 3′UTR) against ALL in Saudi population. Significant protective relationship between the T allele (OR: 0.53 and p = 0.013) and heterozygous genotype TC in codominant (OR: 0.47 and p = 0.024), dominant (OR: 0.48 and p = 0.015) and overdominant (OR: 0.50 and p = 0.033) models. According to AIC value, the dominant model is the expected model that fits the data as it is the lowest among all models. In previous studies, some associations between SNPs in the IL-17A gene and cancer diseases have been reported. Similar to our finding, Bedoui et al. (2018), found that Tunisian individuals carrying the heterozygous genotype of rs3748067 genotype were associated with a reduced risk of colorectal cancer (OR = 0.56, p = 0.003). Wang et al. (2014) reported that individuals sharing the CC genotype or C allele (minor allele) have high risk to develop gastric cancer in Chinese population, while the T allele and TT genotype confer protective effect. However, in two independent meta-analysis after pooling data from various studies, no association between rs3748067 and gastric cancer was validated (Elshazli et al., 2018; Ying et al., 2018). Another study on 62 acute myeloid leukemia patients and 125 healthy controls conducted by Zhu et al., 2015 Another study on 62 acute myeloid leukemia patients and 125 healthy controls conducted by Zhu et al., 2015 found that IL17A genetic polymorphisms were not associated with the risk of this cancer found that IL17A genetic polymorphisms were not associated with the risk of this cancer(Zhu et al., 2015). Contrary to the protective effect abovementioned of the rs3748067 polymorphism, an increased risk to develop cervical cancer was reported for people sharing the TT genotype as reported by Niu et al., 2017 in a study from Shangqiu, China. The disparities these results could be attributed to racial differences, different patient selection criteria, the sample size, and a different SNP detection assay. The functional effect of the rs3748067 could be attributed to its location at the 3′UTR of the IL-17A gene, ant it might play a role in the regulation of the gene expression following their interaction with MicroRNA molecules (Bartel, 2004). The rs3748067 SNP, was reported to probably influence with the IL17A expression by modifying the binding affinity between miRNAs and the target sequence in the 3′UTR regulatory region (Dong et al., 2017).
On the other hands, for the other two SNPs, rs3819025 and rs8193036, no significant associations with any genotype were found with the disease. However, slightly protective effect was found with the minor allele, but mot confirmed for genotypes. In the literature, few studies have investigated these two SNP in relation with cancer diseases. Similar to our results for the rs3819025, no association with breast cancer (Wang et al., 2012), gastric cancer (Qinghai et al., 2014) and cervical cancer (Li et al., 2015) were found in Chinese populations. Colorectal cancer was not also affected by the rs3819025 in Tunisian population as reported by Bedoui et al. (2018). For the rs8193036 (located in the promotor), while no direct associations have been reported with cancer diseases, some studies have linked the mutations with some clinical feature of cancer. In this context, recent study have reported association between rs8193036 with the EGFR mutation status and more advanced stages (III + IV) among lung adenocarcinoma (LUAD) patients, in a Taiwanese population (Lee et al., 2021). Also, this SNP was significantly associated with breast cancer mortality and all-cause mortality among women with greater native American ancestry but not among Hispanic (Slattery et al., 2014). Our analysis of haplotypes associations, revealed potential protective effects of T-G-C (OR = 0.52 and p = 0.025) and C-G-T (OR = 0.45 and p = 0.0089) of the rs8193036, rs3819025, rs3748067 haplotype. These two haplotypes share the rs3819025 G allele and probably TC heterozygous genotype of rs3748067. For IL-17A gene expression different results have been reported. Some study have associated high levels of proteins or mRNA with a better overall outcomes (Chen et al., 2011), but others reported opposite effect (Meng et al., 2012; Song et al., 2022). In our analysis, we did not find significant differences between ALL and healthy control. This is probably due to the low number of included subjects and the heterogeneity between them. More analysis with consideration of clinical selection criteria are recommended.
In conclusion for this study, we suggest that rs3748067 and the haplotypes CTG and TCG may reduce the risk to ALL. These findings could be helpful in the prognostic of ALL, but further large sample studies are needed to confirm these results.
Acknowledgments
Appreciation to the Deputyship for Research & Innovation, Ministry of Education in Saudi Arabia for funding this research work through the project no. (IFKSURG-2-435).
Statement of Ethics
This research was conducted ethically in accordance with the World Medical Association Declaration of Helsinki. The samples were collected from the Department of Medical Oncology, King Khalid Hospital in Riyadh city. All procedures performed in study that involved human participants were in accordance with the Ethics Committee of the Faculty of Medicine, king Saud University, Riyadh, Saudi Arabia and all subjects provided written informed consent.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Extracellular traps released by antimicrobial T H 17 cells contribute to host defense. J. Clin. Invest.. 2021;131(2)
- [Google Scholar]
- Association between Toll-like receptor 4 polymorphism and Acute Lymphoblastic Leukemia susceptibility in Saudi Arabian patients. J. King Saud Univ. – Sci.. 2022;34(4):101985.
- [Google Scholar]
- Evaluation of the relationships between HLA-G 14 bp polymorphism and two acute leukemia in a Saudi population. J. King Saud Univ. – Sci.. 2022;34(6):102139.
- [Google Scholar]
- MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281-297.
- [Google Scholar]
- Association of interleukin-17A polymorphisms with the risk of colorectal cancer: a case-control study. Cytokine. 2018;110:18-23.
- [CrossRef] [Google Scholar]
- Polymorphisms at the IL17A and IL17RA genes are associated with prognosis of papillary thyroid carcinoma. Arch Med Res. 2022;53(2):163-169.
- [CrossRef] [Google Scholar]
- Intratumoral expression of IL-17 and its prognostic role in gastric adenocarcinoma patients. Int. J. Biol. Sci.. 2011;7(1):53-60.
- [CrossRef] [Google Scholar]
- IL-17A in the tumor microenvironment of the human colorectal adenoma–carcinoma sequence. Scandinavian J. Gastroenterol.. 2012;47(11):1304-1312.
- [Google Scholar]
- Role of IL-17A rs2275913 and IL-17F rs763780 polymorphisms in risk of cancer development: an updated meta-analysis. Sci. Rep.. 2016;6(1):20439.
- [CrossRef] [Google Scholar]
- Role of IL-17A rs2275913 and IL-17F rs763780 polymorphisms in risk of cancer development: an updated meta-analysis. Sci. Rep.. 2016;6(1)
- [Google Scholar]
- Single nucleotide polymorphisms and cancer susceptibility. Oncotarget. 2017;8(66):110635-110649.
- [CrossRef] [Google Scholar]
- Associations of functional MicroRNA binding site polymorphisms in IL23/Th17 inflammatory pathway genes with gastric cancer risk. Mediators Inflamm.. 2017;2017:1-11.
- [Google Scholar]
- IL17A gene polymorphism, serum IL17 and total IgE in Egyptian population with chronic HCV and hepatocellular carcinoma. Immunol. Lett.. 2015;168(2):240-245.
- [Google Scholar]
- Genetic polymorphisms of IL-17A rs2275913, rs3748067 and IL-17F rs763780 in gastric cancer risk: evidence from 8124 cases and 9873 controls. Mol. Biol. Rep.. 2018;45(5):1421-1444.
- [Google Scholar]
- Somatic mosaicism: on the road to cancer. Nat. Rev. Cancer. 2016;16(1):43-55.
- [CrossRef] [Google Scholar]
- Interleukin 17–producing CD4+ effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nat. Immunol.. 2005;6(11):1123-1132.
- [Google Scholar]
- Fate mapping of IL-17-producing T cells in inflammatory responses. Nat. Immunol.. 2011;12(3):255-263.
- [Google Scholar]
- Interleukin-17A gene polymorphism is associated with susceptibility to gastric cancer. Int. J. Clin. Experimental Pathol.. 2015;8(6):7378.
- [Google Scholar]
- Functional specialization of Interleukin-17 family members. Immunity. 2011;34(2):149-162.
- [Google Scholar]
- Commensal microbiota promote lung cancer development via γδ T Cells. Cell. 2019;176(5):998-1013.e1016.
- [CrossRef] [Google Scholar]
- Involvement of CCR6/CCL20/IL-17 Axis in NSCLC Disease Progression. PLoS One. 2011;6(9):e24856.
- [Google Scholar]
- Activated monocytes in peritumoral stroma of hepatocellular carcinoma promote expansion of memory T helper 17 cells. Hepatology. 2010;51(1):154-164.
- [Google Scholar]
- Non-Coding variants in cancer: mechanistic insights and clinical potential for personalized medicine. Noncoding RNA. 2021;7(3)
- [CrossRef] [Google Scholar]
- High prevalence of Foxp3 and IL17 in MMR-proficient colorectal carcinomas. Gut. 2008;57(6):772-779.
- [Google Scholar]
- Association between interleukin 17/interleukin 17 receptor gene polymorphisms and papillary thyroid cancer in Korean population. Cytokine. 2015;71(2):283-288.
- [Google Scholar]
- Potential impacts of Interleukin-17A promoter polymorphisms on the EGFR mutation status and progression of non-small cell lung cancer in Taiwan. Genes. 2021;12:427.
- [CrossRef] [Google Scholar]
- Interleukin 17 and peripheral IL-17-expressing T cells are negatively correlated with the overall survival of head and neck cancer patients. Oncotarget. 2018;9(11):9825-9837.
- [Google Scholar]
- Single nucleotide polymorphisms in PLCE1 for cancer risk of different types: a meta-analysis. Front. Oncol.. 2018;8:613.
- [Google Scholar]
- The associations between interleukin-17 single-nucleotide polymorphism and colorectal cancer susceptibility: a systematic review and meta-analysis. World J. Surg. Oncol.. 2022;20(1):116.
- [CrossRef] [Google Scholar]
- Association analysis of IL-17A and IL-17F polymorphisms in Chinese women with cervical cancer. Genet Mol Res. 2015;14(4):12178-12183.
- [CrossRef] [Google Scholar]
- The Accumulation and Prognosis Value of Tumor Infiltrating IL-17 Producing Cells in Esophageal Squamous Cell Carcinoma. PloS one. 2011;6(3):e18219.
- [Google Scholar]
- RETRACTED ARTICLE: Correlation of IL-1F genetic polymorphisms with the risk of colorectal cancer among Chinese populations. Tumor Biol.. 2015;36(2):807-814.
- [Google Scholar]
- Secukinumab inhibition of interleukin-17A in patients with psoriatic arthritis. N. Engl. J. Med.. 2015;373(14):1329-1339.
- [Google Scholar]
- Expression of interleukin-17 and its clinical significance in gastric cancer patients. Med Oncol. 2012;29(5):3024-3028.
- [CrossRef] [Google Scholar]
- IL-17 and IL-17-producing cells in protection versus pathology. Nat. Rev. Immunol. 2022
- [CrossRef] [Google Scholar]
- Generation and regulation of human CD4+ IL-17-producing T cells in ovarian cancer. Proc. Natl. Acad. Sci.. 2008;105(40):15505-15510.
- [Google Scholar]
- Role of IL17A rs2275913 and rs3748067 polymorphisms in the risk cervical cancer. Genet. Mol. Res.. 2017;16(3)
- [Google Scholar]
- Interleukin-17 promotes angiogenesis and tumor growth. J. Am. Soc. Hematol.. 2003;101(7):2620-2627.
- [Google Scholar]
- Involvement of IL17A, IL17F and IL23R polymorphisms in colorectal cancer therapy. PLoS One. 2015;10(6):e0128911.
- [Google Scholar]
- Omrane, I., Marrakchi, R., Baroudi, O., Mezlini, A., Ayari, H., Medimegh, I., Stambouli, N., Kourda, N., Bouzaienne, H., & Uhrhammer, N., 2014. Significant association between interleukin-17A polymorphism and colorectal cancer. 35(7), 6627-6632.
- Early lymphoid development and microenvironmental cues in B-cell acute lymphoblastic leukemia. Arch. Med. Res.. 2012;43(2):89-101.
- [Google Scholar]
- Effect of interleukin-17A and interleukin-17F gene polymorphisms on the risk of gastric cancer in a Chinese population. Gene. 2014;537(2):328-332.
- [CrossRef] [Google Scholar]
- Association of Inflammation-Related and microRNA Gene Expression with Cancer-Specific Mortality of Colon AdenocarcinomaInflammatory Genes, miR-21 and Colon Cancer Prognosis. Clin. Cancer Res.. 2009;15(18):5878-5887.
- [Google Scholar]
- Phenotypic analysis of prostate-infiltrating lymphocytes reveals TH17 and Treg skewing. Clin. Cancer Res.. 2008;14(11):3254-3261.
- [Google Scholar]
- Genetic variants in interleukin genes are associated with breast cancer risk and survival in a genetically admixed population: the Breast Cancer Health Disparities Study. Carcinogenesis. 2014;35(8):1750-1759.
- [CrossRef] [Google Scholar]
- SNPStats: a web tool for the analysis of association studies. Bioinformatics. 2006;22(15):1928-1929.
- [Google Scholar]
- Genetically Predicted Circulating Levels of Cytokines and the Risk of Cancer. Front Immunol. 2022;13:886144
- [CrossRef] [Google Scholar]
- IL-17 family cytokines mediated signaling in the pathogenesis of inflammatory diseases. Cellular Signalling. 2013;25(12):2335-2347.
- [Google Scholar]
- Tumor microenvironments direct the recruitment and expansion of human Th17 cells. J. Immunol.. 2010;184(3):1630-1641.
- [Google Scholar]
- Interleukin-17A modulates circulating tumor cells in tumor draining vein of colorectal cancers and affects MetastasesIL-17A modulates change of CTCs and affects metastases. Clin. Cancer Res.. 2014;20(11):2885-2897.
- [Google Scholar]
- Interleukin 17 is a chief orchestrator of immunity. Nat. Immunol.. 2017;18(6):612-621.
- [Google Scholar]
- Association analysis of IL-17A and IL-17F polymorphisms in Chinese Han women with breast cancer. PLoS One. 2012;7(3):e34400.
- [Google Scholar]
- Elevated levels of IL-17A and IL-35 in plasma and bronchoalveolar lavage fluid are associated with checkpoint inhibitor pneumonitis in patients with non-small cell lung cancer. Oncol. Lett.. 2020;20(1):611-622.
- [CrossRef] [Google Scholar]
- IL-17 gene polymorphism is associated with susceptibility to gastric cancer. Tumour Biol.. 2014;35(10):10025-10030.
- [CrossRef] [Google Scholar]
- The Role of Interleukin 17 in tumour proliferation, angiogenesis, and metastasis. Mediators Inflamm.. 2014;2014:1-12.
- [Google Scholar]
- IL-17 Gene Rs3748067 C>T polymorphism and gastric cancer risk: a meta-analysis. Open Life Sci.. 2018;13:71-76.
- [CrossRef] [Google Scholar]
- The prevalence of Th17 cells in patients with gastric cancer. Biochem. Biophys. Res. Commun.. 2008;374(3):533-537.
- [Google Scholar]
- The role of interleukin-17 in tumor development and progression. J. Exp. Med.. 2020;217(1)
- [Google Scholar]
- Correlation between acute myeloid leukemia and IL-17A, IL-17F, and IL-23R gene polymorphism. Int. J. Clin. Experimental Pathol.. 2015;8(5):5739.
- [Google Scholar]
Appendix A
Supplementary material
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jksus.2022.102493.
Appendix A
Supplementary material
The following are the Supplementary data to this article:







